NEET Previous Year Questions(2016-24): Molecular Basis of Inheritance | Biology Class 12 PDF Download
2024
Q1: A transcription unit in DNA is defined primarily by the three regions in DNA and these are with respect to upstream and down stream end; (NEET 2024)
(a) Repressor, Operator gene, Structural gene
(b) Structural gene, Transposons, Operator gene
(c) Inducer, Repressor, Structural gene
(d) Promotor, Structural gene, Terminator
Ans: (d)
A transcription unit in DNA is critical for the process of transcription, wherein a particular segment of DNA is copied into RNA (especially mRNA) by the enzyme RNA polymerase. This unit is composed of sequences that include both coding regions, which are directly transcribed into RNA, and regulatory regions, which ensure that transcription is initiated and terminated at the correct locations on the DNA.
The correct answer is: Option D: Promotor, Structural gene, Terminator
Here's a detailed explanation of each component:
Promoter: The promoter is a sequence in DNA that signals the RNA polymerase to start transcription. It is located at the upstream end (5' end) of the gene. Promoters are essential for transcription initiation and are typically found just before the genes they regulate.
Structural gene: This region of the transcription unit is actually expressed or translated into protein (or functional RNA), depending on the kind of gene. These genes contain the functional sequences that are copied during the transcription process.
Terminator: The terminator is found at the downstream end (3' end) of the transcription unit and includes sequences that signal the RNA polymerase enzyme to stop transcription. This ensures that the newly synthesized RNA contains only the necessary genetic message.
The other options contain components that do not accurately define the typical structure of a transcription unit:
Option A mixes regulatory proteins and DNA regions, which does not accurately represent the structural components of a transcription unit.
Option B includes "transposons" which are genetic elements that can move around within the genomes but are not typically part of the transcription unit.
Option C again refers to regulatory proteins (inducer and repressor) along with structural genes, confusing the functions of proteins and DNA regions.
Therefore, Option D correctly represents the standard components of a transcription unit in the context of gene transcription in DNA.
Q2: Match List I with List II (NEET 2024)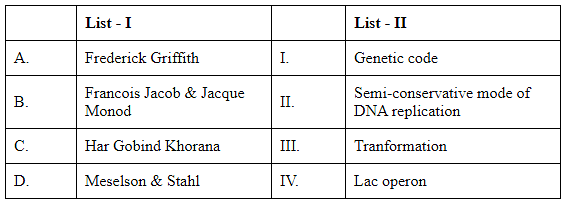
Choose the correct answer from the options given below:
(a) A-III, B-II, C-I, D-IV
(b) A-III, B-IV, C-I, D-II
(c) A-II, B-III, C-IV, D-I
(d) A-IV, B-II, C-II, D-III
Ans: (b)
To solve this matching question, let's discuss each scientist(s) and their contribution:
A. Frederick Griffith: Known for discovering the "transforming principle," which showed that a substance from dead bacteria could genetically transform living bacteria. This process is called transformation. The correct match for Frederick Griffith is III. Transformation.
B. Francois Jacob & Jacques Monod: They are famous for their work on the lac operon, a set of genes involved in lactose metabolism in bacteria. Their study elucidated how genes are regulated and expressed in cells. The correct match for Francois Jacob and Jacques Monod is IV. Lac operon.
C. Har Gobind Khorana: Known for his research on the genetic code and its role in protein synthesis. Khorana was one of the scientists who elucidated how the sequence of nucleotides in nucleic acids is translated into protein sequences. The correct match for Har Gobind Khorana is I. Genetic code.
D. Meselson & Stahl: Famous for their experiment confirming the semi-conservative mechanism of DNA replication, where each new DNA molecule consists of one old strand and one newly synthesized strand. The correct match for Meselson and Stahl is II. Semi-conservative mode of DNA replication.
Comparing this information with the options provided:
Option A: A-III, B-II, C-I, D-IV (Incorrect: B does not match II)
Option B: A-III, B-IV, C-I, D-II (Correct: All matches are accurate)
Option C: A-II, B-III, C-IV, D-I (Incorrect: A, B, C, and D do not match correctly)
Option D: A-IV, B-II, C-II, D-III (Incorrect: A, C, and D do not match correctly)
Therefore, the correct answer is Option B.
Q3: Which of the following statement is correct regarding the process of replication in E.coli? (NEET 2024)
(a) 'The DNA dependent DNA polymerase catalyses polymerization in one direction that is 3' → 5'
(b) The DNA dependent RNA polymerase catalyses polymerization in one direction, that is 5' → 3'
(c) The DNA dependent DNA polymerase catalyses polymerization in 5' → 3' as well as 3' → 5' direction
(d) The DNA dependent DNA polymerase catalyses polymerization in 5' → 3' direction.
Ans: (d)
The correct statement regarding the process of replication in E.coli is found in Option D: "The DNA dependent DNA polymerase catalyses polymerization in 5' → 3' direction".
To elaborate, replication in E.coli involves the synthesis of new DNA strands from a DNA template. This synthesis is catalyzed by an enzyme known as DNA polymerase. DNA polymerases are key enzymes that add nucleotides to the growing DNA strand during replication. However, the key characteristic of these enzymes is their directionality. DNA polymerases can only add nucleotides to the 3' end of the DNA strand, thereby synthesizing new DNA in the direction of 5' → 3'.
This directional limitation is due to the structure of deoxyribonucleotide triphosphates (dNTPs) that are used as substrates by the enzyme. Each dNTP has a 5' phosphate group, a 3' hydroxyl group, and a nitrogenous base. The formation of DNA strands occurs via the formation of phosphodiester bonds between the 3' hydroxyl group of one nucleotide and the phosphate group at the 5' position of the next nucleotide. Therefore, nucleotides can only be added to the 3' end of the growing strand.
The other options are incorrect for the following reasons:
Option A incorrectly states the direction of synthesis as which is impossible with the mechanics of DNA polymerases.
Option B refers to DNA dependent RNA polymerase. This enzyme is indeed involved in transcription (synthesizing RNA from DNA), not replication, and synthesizes RNA in the direction.
Option C incorrectly states that DNA polymerase can synthesize in both directions, which contradicts the inherent directional nature of this enzyme
Therefore, understanding the precise enzyme mechanics and functionality helps in recognizing that Option D is the correct description of how DNA dependent DNA polymerase facilitates DNA replication in E.coli.
Q4: Which one is the correct product of DNA dependent RNA polymerase to the given template? (NEET 2024)
3'TACATGGCAAATATCCATTCA5'
(a) 5'AUGUACCGUUUAUAGGUAAGU3'
(b) 5'AUGUAAAGUUUAUAGGUAAGU3'
(c) 5'AUGUACCGUUUAUAGGGAAGU3'
(d) 5'ATGTACCGTTTATAGGTAAGT3'
Ans: (a)
DNA-dependent RNA polymerase is an enzyme responsible for transcribing DNA into RNA. During transcription, the RNA polymerase reads the template strand of DNA and synthesizes a complementary RNA strand. A key point in this process is that RNA polymerase builds RNA by replacing thymine (T) with uracil (U).
The provided DNA template is: 3'TACATGGCAAATATCCATTCA5'
To find the correct RNA sequence, we need to identify the complementary base for each base in the template strand while considering RNA bases. Remember, in RNA:
A (Adenine) pairs with U (Uracil)
T (Thymine) pairs with A (Adenine)
C (Cytosine) pairs with G (Guanine)
G (Guanine) pairs with C (Cytosine)
The complementary RNA sequence to the DNA template is generated as follows:
3'T --> 5'A
3'A --> 5'U
3'C --> 5'G
3'G --> 5'C
3'T --> 5'A
3'A --> 5'U
3'T --> 5'A
3'G --> 5'C
3'G --> 5'C
3'C --> 5'G
3'A --> 5'U
3'A --> 5'U
3'T --> 5'A
3'A --> 5'U
3'T --> 5'A
3'C --> 5'G
3'C --> 5'G
3'A --> 5'U
3'T --> 5'A
3'T --> 5'A
3'C --> 5'G'
This constructs the RNA sequence: 5'AUGUACCGUUUAUAGGUAAGU3'
Thus, Option A correctly represents the RNA sequence transcribed by DNA-dependent RNA polymerase from the given DNA template:
Option A: 5'AUGUACCGUUUAUAGGUAAGU3'
Q5: Match List I with List II: (NEET 2024)
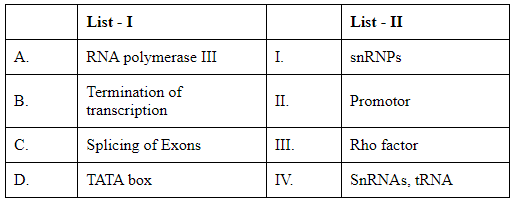 Choose the correct answer from the options given below :
Choose the correct answer from the options given below :
(a) A-II, B-IV, C-I, D-III
(b) A-III, B-II, C-IV, D-I
(c) A-III, B-IV, C-I, D-II
(d) A-IV, B-III, C-I, D-II
Ans: (d)
The goal is to correctly match the terms in List I with the descriptions in List II. Let's analyze and match each term from List I with its appropriate partner in List II:
List I:
A. RNA polymerase III: This enzyme is primarily responsible for transcribing DNA to synthesize tRNA, 5S rRNA, and other small RNAs.
B. Termination of transcription: In prokaryotes, specific termination factors such as the Rho factor are involved in stopping transcription. In eukaryotes, different mechanisms and sequences are used.
C. Splicing of Exons: The process involving the removal of introns and joining of exons during mRNA processing. Small nuclear ribonucleoproteins (snRNPs) are critical components in this process.
D. TATA box: A DNA sequence within the promoter region, which is crucial for forming the transcription initiation complex.
List II:
I. snRNPs: Small nuclear ribonucleoproteins involved in mRNA splicing.
II. Promoter: A region of DNA that initiates transcription of a particular gene, typically containing sequences like the TATA box.
III. Rho factor: A protein essential for terminating transcription in prokaryotes.
IV. SnRNAs, tRNA: Molecules transcribed primarily by RNA polymerase III.
Matching these descriptions:
A (RNA polymerase III) matches with IV (SnRNAs, tRNA).
B (Termination of transcription) matches with III (Rho factor).
C (Splicing of Exons) matches with I (snRNPs).
D (TATA box) matches with II (Promoter).
Therefore, the correct matches according to options listed are: Option D: A-IV, B-III, C-I, D-II
Q6: The lactose present in the growth medium of bacteria is transported to the cell by the action of (NEET 2024)
(a) Beta-galactosidase
(b) Acetylase
(c) Permease
(d) Polymerase
Ans: (c)
The correct answer is Option C: Permease.
Lactose, a disaccharide sugar composed of glucose and galactose, needs particular mechanisms to be transported into bacterial cells for metabolism. Among the options provided, permease is the protein responsible for this function. Specifically, in the case of bacterial cells such as Escherichia coli, the lactose permease enzyme plays a crucial role.
Lactose permease is encoded by the lacY gene, which is part of the lac operon. The lac operon is a famous example of gene regulation in bacteria. When lactose is present outside the cell, lactose permease facilitates its transport into the cell across the cell membrane. Once inside, lactose can be utilized as a source of energy and carbon.
Once lactose is inside the bacterial cell:
It is broken down by the enzyme β-galactosidase into glucose and galactose. This enzyme is encoded by the lacZ gene, which is a different component of the lac operon.
We can rule out the other options because:
Beta-galactosidase (Option A) is involved in the hydrolysis of lactose into glucose and galactose but not in its transport.
Acetylase (Option B) refers to enzymes involved in the addition of acetyl groups to substrates, unrelated to sugar transport.
Polymerase (Option D) are enzymes that synthesize RNA and DNA, thus also unrelated to transporting sugars like lactose.
Thus, to facilitate the transport of lactose across the cell membrane into the bacterial cell, lactose permease (Option C) is required, making it the correct choice.
2023
Q1: Unequivocal proof that DNA is the genetic material was first proposed by (NEET 2023)
(a) Frederick Griffith
(b) Alfred Hershey and Martha Chase
(c) Avery, Macleoid and McCarthy
(d) Wilkins and Franklin
Ans: (b)
- The first unequivocal proof that DNA is the genetic material came from the experiments conducted by Alfred Hershey and Martha Chase in 1952. They used bacteriophages (viruses that infect bacteria) and radioactively labeled the protein and DNA of the phage separately in two different sets of experiments. They found that it was the DNA, not the protein, of the phage that was injected into the bacteria and carried the genetic information necessary for the production of new phage particles.
- Avery, Macleoid and McCarty gave the biochemical characterisation of Transforming Principle.
- The transformation experiments by using Pneumococcus was conducted by Frederick Griffith.
- Wilkins and Franklin produced X-ray diffraction data of DNA.
So, the correct answer is : Option B : Alfred Hershey and Martha Chase.
Q2: What is the role of RNA polymerase III in the process of transcription in Eukaryotes? (NEET 2023)
(a) Transcription of rRNAs (28S, 18S and 5.8S)
(b) Transcription of tRNA, 5S rRNA and snRNA
(c) Transcription of precursor of mRNA
(d) Transcription of only snRNAs
Ans: (b)
- In eukaryotes there are three major types of RNA polymerases.
- RNA polymerase I transcribes : 5.8S, 18S, 28S rRNAs
- RNA polymerase II transcribes : hnRNAs (precurssor of mRNA)
- RNA polymerase III transcribes : tRNAs, ScRNA, 5S rRNA and snRNA
Q3: Expressed Sequence Tags (ESTs) refers to (NEET 2023)
(a) All genes that are expressed as RNA.
(b) All genes that are expressed as proteins.
(c) All genes whether expressed or unexpressed.
(d) Certain important expressed genes.
Ans: (a)
Expressed Sequence Tags (ESTs) are short sub-sequences of a cDNA sequence. They may identify expressed genes, so they are derived from mRNA which is transcribed from expressed genes. They serve as a kind of "tag" or marker for identifying the gene from which it was transcribed. Therefore, ESTs represent genes that are expressed as RNA. The other options listed do not accurately describe what ESTs are.
Q4: Upon exposure to UV radiation, DNA stained with ethidium bromide will show (NEET 2023)
(a) Bright blue colour
(b) Bright yellow colour
(c) Bright orange colour
(d) Bright red colour
Ans: (c)
Option (C) is the correct answer because in recombinant DNA technology the separated DNA fragments can be visualised only after staining the DNA with a substance known as ethidium bromide followed by exposure to U.V. radiation. You can see bright orange coloured bands of DNA in an ethidium bromide stained gel exposed to U.V. light.
Q5: Match List I with List II. (NEET 2023)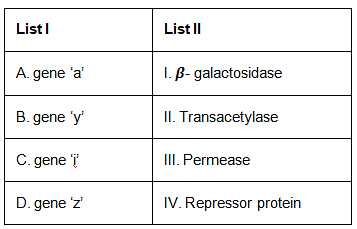
Choose the correct answer from the options given below:
(a) A-II, B-I, C-IV, D-III
(b) A-II, B-III, C-IV, D-I
(c) A-III, B-IV, C-I, D-II
(d) A-III, B-I, C-IV, D-II
Ans: (b)
The question relates to the lac operon model in E. coli, which is a well-studied example of gene regulation. In this model :
- Gene 'z' codes for beta-galactosidase (breaks down lactose into glucose and galactose)
- Gene 'y' codes for permease (transports lactose into the cell)
- Gene 'a' codes for transacetylase (may help in lactose metabolism, but its exact role is unclear)
- Gene 'i' codes for the repressor protein (binds to the operator to prevent transcription)
So, matching the List I (genes) with List II (proteins they code for) :
- A (Gene 'a') matches with II (Transacetylase)
- B (Gene 'y') matches with III (Permease)
- C (Gene 'i') matches with IV (Repressor protein)
- D (Gene 'z') matches with I (Beta-galactosidase)
Therefore, the correct option is : Option A : A-II, B-III, C-IV, D-I
Q6: Given below are two statements: (NEET 2023)
Statement I: In prokaryotes, the positively charged DNA is held with some negatively charged proteins in a region called nucleoid.
Statement II: In eukaryotes, the negatively charged DNA is wrapped around the positively charged histone octamer to form nucleosome.
In the light of the above statements, choose the correct answer from the options given below:
(a) Both Statement I and Statement II are true.
(b) Both Statement I and Statement II are false.
(c)Statement I is correct but Statement II is false.
(d) Statement I is incorrect but Statement II is true.
Ans: (d)
- In prokaryotes, the negatively charged DNA is held with some positively charged proteins in a region termed as nucleoid.
- In eukaryotes, the negatively charged DNA is wrapped around the positively charged histone octamer to form a structure called nucleosome.
Q7: Which one of the following is the sequence on corresponding coding strand, if the sequence on mRNA formed is as follows 5’AUCGAUCGAUCGAUCGAUCGAUCG AUCG 3’? (NEET 2023)
(a) 5’ UAGCUAGCUAGCUAGCUAGCUAGCUAGC 3’
(b) 3’ UAGCUAGCUAGCUAGCUAGCUAGCUAGC 5’
(c) 5’ ATCGATCGATCGATCGATCGATCGATCG 3’
(d) 3’ ATCGATCGATCGATCGATCGATCGATCG 5’
Ans: (c)
The sequence on the mRNA is 5’AUCGAUCGAUCGAUCGAUCGAUCG AUCG 3’. The mRNA is synthesized from the template strand of DNA by complementary base pairing, following these rules: A pairs with U, U pairs with A, C pairs with G, and G pairs with C. The coding strand of DNA has the same sequence as the mRNA, except with T instead of U.
To find the sequence on the corresponding coding strand, we can use these rules to convert the mRNA sequence back to DNA :
5’ AUCGAUCGAUCGAUCGAUCGAUCG AUCG 3’ (mRNA)
5’ ATCGATCGATCGATCGATCGATCG ATCG 3’ (coding strand DNA)

2022
Q1: The process of translation of mRNA to proteins begins as soon as: (NEET 2022)
(a) The larger subunit of ribosome encounters mRNA
(b) Both the subunits join together to bind with mRNA
(c) The tRNA is activated and the larger subunit of ribosome encounters mRNA
(d) The small subunit of ribosome encounters mRNA
Ans: (d)
When the small subunit of ribosome encounters an mRNA, the process of translation of the mRNA to protein begins. This process is followed by the binding of bigger/larger subunit.
t-RNA is activated by the addition of amino acid prior to the attachment or ribosome, in the first phase.
Q2: Read the following statements and choose the set of correct statements: (NEET 2022)
(A) Euchromatin is loosely packed chromatin
(B) Heterochromatin is transcriptionally active
(C) Histone octomer is wrapped by negatively charged DNA in nucleosome
(D) Histones are rich in lysine and arginine
(E) A typical nucleosome contains 400 bp of DNA helix
Choose the correct answer from the options given below:
(a) (A), (C), (D) Only
(b) (B), (E) Only
(c) (A), (C), (E) Only
(d) (B), (D), (E) Only
Ans: (a)
- Heterochromatin is transcriptionally inactive. A typical nucleosome contains 200 bp of DNA helix.
- Euchromatin is the loosely packed chromatin region.
- The negatively charged DNA is wrapped around the positively charged histone octamer to form a structure called nucleosome. Histones are rich in basic amino acid residues lysine and arginine.
Q3: DNA Polymorphism forms the basis of: (NEET 2022)
(a) DNA finger printing
(b) Both genetic mapping and DNA finger printing
(c) Translation
(d) Genetic mapping
Ans: (b)
Polymorphism in DNA sequence is the basis of genetic mapping of human genome as well as of DNA finger printing.
Q4: If the length of a DNA molecule is 1.1 metres, what will be the approximate number of base pairs? (NEET 2022)
(a) 6.6 x 109 bp
(b) 3.3 x 106 bp
(c) 6.6 x 106 bp
(d) 3.3 x 109 bp
Ans: (d)
The two chains of DNA are coiled in a right-handed helicle fashion. The pitch of the helix is 3.4 nm and has a diameter of 2 nm. In the B-form of DNA, the distance between a bp in a helix is approximately 0.34 nm which means there are about 10 nucleotides present in a complete turn.
The total length of DNA molecule in nm = 1.1 × 109 nm.
The approximate number of base pairs = 1.1 × 109/ 0.34
= 3.3 × 109
Q5: Transposons can be used during which one of the following? (NEET 2022)
(a) Polymerase Chain Reaction
(b) Gene Silencing
(c) Autoradiography
(d) Gene sequencing
Ans: (b)
- Option (b) is the correct answer as the source of the complementary RNA for RNAi could be mobile genetic elements (transposons) that replicate via an RNA intermediate.
- Option (c) is incorrect as autoradiography usually follows hybridisation.
- Option (a) is incorrect because polymerase chain reaction is used to make copies of the DNA sample and does not need transposons.
- Option (d) is incorrect because transposons are not required during gene sequencing.
Q6: If a geneticist uses the blind approach for sequencing the whole genome of an organism, followed by assignment of function to different segments, the methodology adopted by him is called as : (NEET 2022)
(a) Sequence annotation
(b) Gene mapping
(c) Expressed sequence tags
(d) Bioinformatics
Ans: (a)
Sequencing the whole set of genome that contained all the coding and non-coding sequences and later assigning different regions in the sequence with functions is called sequence annotation.
Q7: In an E. coli strain, i gene gets mutated, and its product cannot bind the inducer molecule. If the growth medium is provided with lactose, what will be the outcome? (NEET 2022)
(a) a genes will be transcribed
(b) a genes will not be translated
(c) RNA polymerase will bind the promoter region
(d) Only gene will get transcribed
Ans: (a)
- As the product of 'i' gene binds with the operator region and blocks the transcription and translation of z, y and a genes.
- It's product is prevented from binding to the operator by attaching it with the inducer. As the inducer can not no more capable of binding with the repressor, thus, in all the cases, operator always gets attached with the repressor thereby preventing the transcription and transmission of z, y and a.
- Even in the presence of lactose, transcription and translation of z, y and a would not occur.
Q8: Ten E.coli cells with 15N - dsDNA are incubated in medium containing 14N nucleotide. After 60 minutes, how many E.coli cells will have DNA totally free from 15N?
(a) 20 cells
(b) 40 cells
(c) 60 cells
(d) 80 cells (NEET 2022)
Ans: (c)
From 10 parent E.coli cells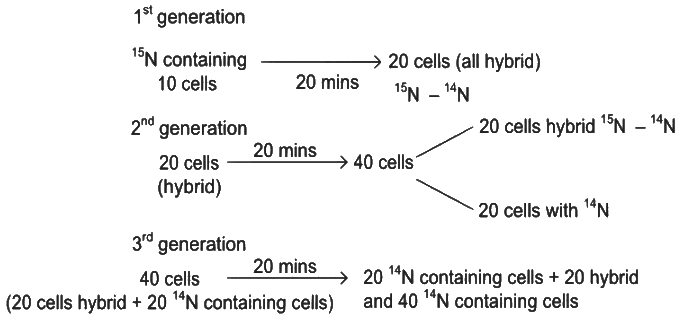
Therefore, after 60 minutes, 60 E.coli cells will have DNA totally free from 15N.
Q9: In lac operon, z gene codes for (NEET 2022 Phase 2)
(a) Transacetylase
(b) β-galactosidase
(c) Permease
(d) Repressor
Ans: (b)
In lac operon, z gene codes for β-galactosidase.
Transacetylase, permease and repressor protein are coded by genes 'a', 'y' and 'i' respectively.
Q10: Given below are two statements (NEET 2022 Phase 2)
Statement I : DNA polymerases catalyse polymerisation only in one direction, that is 5' → 3'.
Statement II : During replication of DNA, on one strand the replication is continuous while on other strand it is discontinuous.
In the light of the above statements, choose the correct answer from the options given below:
(a) Statement I is incorrect but Statement II is correct
(b) Both Statement I and Statement II are correct
(c) Both Statement I and Statement II are incorrect
(d) Statement I is correct but Statement II is incorrect
Ans: (b)
The DNA-dependent DNA polymerases catalyse polymerisation only in one direction, that is 5' → 3'. This creates some additional complications at the replicating fork. Consequently, on one strand (the template with polarity 3' → 5'), the replication is continuous, while on the other (the template with polarity 5' → 3'), it is discontinuous.
Q11: Match List - I with List - II. (NEET 2022 Phase 2)
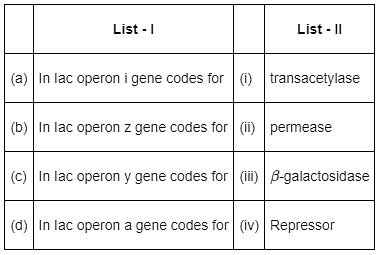
Choose the correct answer from the options given below
(a) (a) - (iii), (b) - (i), (c) - (iv), (d) - (ii)
(b) (a) - (iii), (b) - (ii), (c) - (i), (d) - (iv)
(c) (a) - (iv), (b) - (iii), (c) - (ii), (d) - (i)
(d) (a) - (iv), (b) - (i), (c) - (iii), (d) - (ii)
Ans: (c)
In Iac operon,
- The i gene codes for repressor protein.
- The z gene codes for β-galactosidase.
- The y gene codes for permease and the a gene codes for transacetylase.
Q12: Match List-I with List-II: (NEET 2022 Phase 2)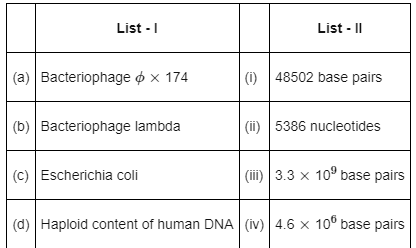
Choose the correct answer from the options given below:
(a) (a)-(i), (b)-(ii), (c)-(iv), (d)-(iii)
(b) (a)-(i), (b)-(ii), (c)-(iii), (d)-(iv)
(c) (a)-(ii), (b)-(iv), (c)-(i), (d)-(iii)
(d) (a)-(ii), (b)-(i), (c)-(iv), (d)-(iii)
Ans: (d)
Genetic material of –
- Bacteriophage ∅ × 174 contains 5386 nucleotides
- Bacteriophage lambda contains 48502 base pairs
- Escherichia coli contains 4.6 × 106 base pairs
- Haploid content of human DNA contains 3.3 × 109 base pairs
Q13: Against the codon 5' UAC 3', what would be the sequence of anticodon on tRNA? (NEET 2022 Phase 2)
(a) 5' GUA 3'
(b) 5' AUG 3'
(c) 5' ATG 3'
(d) 5' GTA 3'
Ans: (b)
- In the process of translation, the mRNA codon pairs with the tRNA anticodon to bring the correct amino acid to the growing polypeptide chain. The pairing between the codon and anticodon follows the complementary base-pairing rules, with adenine (A) pairing with uracil (U), and guanine (G) pairing with cytosine (C).
- In this case, the mRNA codon is given as 5' UAC 3'. To determine the tRNA anticodon sequence, we must follow the base-pairing rules:
- The U (uracil) in the codon pairs with A (adenine) in the anticodon.
- The A (adenine) in the codon pairs with U (uracil) in the anticodon.
- The C (cytosine) in the codon pairs with G (guanine) in the anticodon.
- Therefore, the anticodon sequence on the tRNA would be 5' AUG 3'.
Q14: If A and C make 30% and 20% of DNA, respectively, what will be the percentage composition of T and G?
(a) T : 20%, G : 20%
(b) T : 20%, G : 30%
(c) T : 30%, G : 20%
(d) T : 30%, G : 30% (NEET 2022 Phase 2)
Ans: (c)
According to Chargaff’s rule, amount of Adenine (A) is equal to thymine and amount of cytosine (c) will be equal to that of Guanine.
So,
A = T = 30% + 30% = 60%
C = G = 20% + 20% = 40%
So, T and G will be 30% and 20% respectively.
2021
Q1: Complete the flow chart on central dogma (NEET 2021)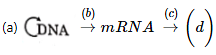
(a) (a) - Replication; (b) - Transcription;
(c) - Translation; (d) - Protein
(b) (a) - Transduction; (b) - Translation;
(c) - Replication; (d) - Protein
(c) (a) - Replication; (b) - Transcription;
(c) - Transduction; (d) - Protein
(d) (a) - Transcription; (b) - Replication;
(c) - Transcription; (d) - Transduction
Ans: (a)
- Formation of DNA from DNA is replication.
- Formation of mRNA from DNA is called Transcription.
- Formation of protein from mRNA is called Translation.
So,
(a) is Replication
(b) is Transcription
(c) is Translation
(d) is Protein
- Transduction is transfer of genetic material from one bacterium to another with the help of virus or a bacteriophage.
Q2: What is the role of RNA polymerase III in the process of transcription in eukaryotes? (NEET 2021)
(a) Transcribes precursor of mRNA
(b) Transcribes only snRNAs
(c) Transcribes rRNAs (28S, 18S, and 5.8S)
(d) Transcribes tRNA, 5s rRNA and snRNA
Ans: (d)
- RNA polymerase III transcribes tRNA, ScRNA, 5S rRNA and snRNA.
- RNA polymerase I transcribes 5.8S, 18S and 28S rRNA.
- RNA polymerase II transcribes hnRNA which is precursor of mRNA
Q3: DNA strands on a gel stained with ethidium bromide when viewed under UV radiation, appear as
(a) Bright blue bands
(b) Yellow bands
(c) Bright orange bands
(d) Dark red bands (NEET 2021)
Ans: (d)
To make the DNA visible in the gel, ethidium bromide is added to the gel solution and the buffer. This positively charged polycyclic aromatic compound binds to DNA by inserting itself between the basepairs (intercalation). The DNA fragments when exposed to ultraviolet light appear as orange colour bands, due to the large increase in fluorescence of the ethidium bromide upon binding to the DNA.
Q4: DNA fingerprinting involves identifying differences in some specific regions in DNA sequence, called as (NEET 2021)
(a) Single nucleotides
(b) Polymorphic DNA
(c) Satellite DNA
(d) Repetitive DNA
Ans: (d)
- DNA fingerprinting involves identifying differences in some specific regions in DNA sequence called as repetitive DNA.
- The basis of DNA fingerprinting is VNTR (a satellite DNA as probe that show very high degree of polymorphism)
- Polymorphism is the variation at genetic level. Allelic sequence variation has traditionally been described as a DNA polymorphism.
Q5: Identify the correct statement. (NEET 2021)
(a) The coding strand in a transcription unit is copied to an mRNA.
(b) Split gene arrangement is characteristic of prokaryotes.
(c) In capping, methylguanosine triphosphate is added to the 3' end of hnRNA.
(d) RNA polymerase binds with the Rho factor to terminate the process of transcription in bacteria.
Ans: (d)
- Split gene arrangement is characteristic of eukaryotes.
- In capping 5-methyl guanosine triphosphate is added at 5' end of hnRNA.
- At 3' end poly-A tail is added.
- The non coding or template strand is copied to an mRNA. RNA polymerase associate with ρ factor (Rho factor) and it alters the specificity of the RNA polymerase to terminate the processes.
Q6: If Adenine makes 30% of the DNA molecule, what will be the percentage of Thymine, Guanine and Cytosine in it? (NEET 2021)
(a) T:30 ; G:20 ; C:20
(b) T:20 ; G:25 ; C:25
(c) T:20 ; G:30 ; C:20
(d) T:20 ; G:20 ; C:30
Ans: (a)
Chargaff rule - In DNA there is always equality in quantity between the bases A and T and between the bases G and C.
According to Chargaff's rule, for a double stranded DNA,
[A] = [T],
∵ [A] = 30%, ⇒ [T] = 30%
Since [C] = [G]
∴ 100 − [A + T]
= 100 − [30 + 30]
= 100 − 60 = 40%
and C = G = 20% each
∴ [A] = 30%
[T] = 30%
[G] = 20%
[C] = 20%
Q7: Which of the following RNAs is not required for the synthesis of protein? (NEET 2021)
(a) rRNA
(b) siRNA
(c) mRNA
(d) tRNA
Ans: (b)
- siRNA mainly protect the cell from exogenous mRNA attacks. It degrades the growing mRNA and stop gene expression. It is highly specific and reduces the synthesis of particular proteins by reducing the translation of specific messenger RNAs. Hence, siRNA is not required for protein synthesis but is used to reduce its synthesis.
- mRNA is messenger RNA that carries genetic information provided by DNA.
- tRNA carries amino acids to the mRNA during translation.
- rRNA is structural RNA that forms ribosomes which are involved in translation.
Q8: Which is the "Only enzyme" that has "Capability to catalyze Initiation, Elongation, and Termination in the process of transcription in prokaryotes? (NEET 2021)
(a) DNA Ligase
(b) DNase
(c) DNA dependent DNA polymerase
(d) DNA dependent RNA polymerase
Ans: (d)
Prokaryotes utilize one RNA polymerase for transcription of all types of RNA. The enzyme RNA polymerase is needed for RNA formation from DNA, i.e. DNA dependent RNA polymerase. It occurs in the cytoplasm of prokaryotic cells. RNA polymerase is the only enzyme which, has the capability to catalyse all initiation, elongation and termination in prokaryotes.
Q9: Which one of the following statements about Histones is wrong? (NEET 2021)
(a) Histones are rich in amino acids - Lysine and Arginine.
(b) Histones carry a positive charge in the side chain.
(c) Histones are organized to form a unit of 8 molecules.
(d) The pH of histones is slightly acidic.
Ans: (d)
- Histones are rich in basic amino acids residue lysine and arginine with charged side chain.
- There are five types of histone proteins i.e. H1, H2A, H2B, H3 and H4. Four of them occur in pairs to produce a unit of 8 molecules (histone octamer).
- The pH of histones is basic.
Q10: Statement I: The codon 'AUG codes for methionine and phenylalanine.
Statement II: AAA' and 'AAG are both codons that code for the amino acid lysine.
In the light of the above statements, choose the correct answer from the options given below. (NEET 2021)
(a) Statement I is correct but Statement II is false
(b) Statement I is incorrect but Statement II is true
(c) Both Statement I and Statement II are true
(d) Both Statement I and Statement II are false
Ans: (b)
- The codon AUG only codes for methionine. As the codons are universal. From bacteria to mammals AUG only codes for methionine.
∴ Statement I is false.
- Some amino acids are coded by more than one codon, hence the code is degenerate. AAA and AAG both codons code for the amino acid lysine.
∴ Statement II is true.
2020
Q1: If the distance between two consecutive base pairs is 0.34 nm and the total number of base pairs of a DNA double helix in a typical mammalian cell is 6.6 × 109 dp, then the length of the DNA is approximately. (NEET 2020)
(a) 2.2 meters
(b) 2.7 meters
(c) 2.0 meters
(d) 2.5 meters
Ans: (a)
The diploid content of human genome is 6.6 x 109 base pairs. The distance between two consecutive base pairs is 0.34 nm (0.34 x 10-9 m), so the length of DNA double helix in a typical mammalian cell is around 6.6 x 10-9 bp x 0.34 x 10-9m/bp = 2.2 metres.
Q2: The first phase of translation is: (NEET 2020)
(a) Aminoacylation of tRNA
(b) Recognition of an anti-codon
(c) Binding of mRNA to ribosome
(d) Recognition of DNA molecule
Ans: (a)
The first phase of translation involves activation of amino acid in the presence of ATP and linked to their cognate tRNA - a process commonly called as charging of tRNA or aminoacylation of tRNA.
Q3: Name the enzyme that facilitates opening of DNA helix during transcription. (NEET 2020)
(a) DNA polymerase
(b) RNA polymerase
(c) DNA ligase
(d) DNA helicase
Ans: (b)
- The enzyme that facilitates opening of the DNA helix during transcription is RNA polymerase.
- RNA polymerase is responsible for the synthesis of RNA from a DNA template during transcription. It binds to the DNA at a specific site called the promoter region and then unwinds the DNA helix to expose the template strand. This opening of the DNA helix is facilitated by the activity of RNA polymerase, which breaks the hydrogen bonds between the base pairs of the DNA.
- Once the DNA is unwound, RNA polymerase can begin the synthesis of RNA by adding complementary RNA nucleotides to the template strand. The RNA polymerase moves along the DNA strand, opening the helix as it goes, and synthesizing a new RNA molecule.
Therefore, option B, "RNA polymerase," is the correct answer.
Q4: Which of the following statements is correct?
(a) Adenine does not pair with thymine
(b) Adenine pairs with thymine through one H-bond
(c) Adenine pairs with thymine through two H-bonds
(d) Adenine pairs with thymine through three H-bonds (NEET 2020)
Ans: (c)
Based on the observation of Erwin Chargaff that for a double stranded DNA, the ratios between Adenine and Thymine and Guanine and Cytosine are constant and equals one. Adenine pairs with thymine through two H-bonds i.e., A = T and guanine pairs with cytosine with three H-bonds.
2019
Q1: Purines found both in DNA and RNA are (NEET 2019)
(a) Cytosine and thymine
(b) Adenine and thymine
(c) Adenine and guanine
(d) Guanine and cytosine
Ans: (c)
Adenine and guanine are purines which are common to both DNA and RNA.
Q2: Expressed Sequence Tags (ESTs) refers to (NEET 2019)
(a) Novel DNA sequences
(b) Genes expressed as RNA
(c) Polypeptide expression
(d) DNA polymorphism
Ans: (b)
Expressed Sequence Tags (ESTs) are genes that are expressed as RNA. It is used in sequencing of human genome.
Q3: Match the following genes of the Lac operon with their respective products. (NEET 2019)
| (A) I gene | (i) β - galactosidase |
| (B) z gene | (ii) Permease |
| (C) a gene | (iii) Repressor |
| (D) y gene | (iv) Transacetylase |
Select the correct option.
| (A) | (B) | (C) | (D) | |
| (a) | (iii) | (iv) | (i) | (ii) |
| (b) | (i) | (iii) | (ii) | (iv) |
| (c) | (iii) | (i) | (ii) | (iv) |
| (d) | (iii) | (i) | (iv) | (ii) |
Ans: (d)
In lac operon
- I gene → Repressor
- z gene → β-galactosidase
- y gene → Permease
- a gene → Transacetylase
Q4:The shorter and longer arms of a submetacentric chromosome are referred to as:
(a) q-arm and p-arm respectively
(b) m-arm and n-arm respectively
(c) s-arm and l-arm respectively
(d) p-arm and q-arm respectively (NEET 2019)
Ans: (d)
The shorter arm of a sub-metacentric chromosome is called as the 'p' - arm and the longer arm is called as a 'q' - arm.
Q5: Which of the following features of genetic code does allow bacteria to produce human insulin by recombinant DNA technology? (NEET 2019)
(a) Genetic code is specific.
(b) Genetic code is not ambiguous.
(c) Genetic code is redundant.
(d) Genetic code is nearly universal.
Ans: (d)
In recombinant DNA technology, a bacterium is able to produce human insulin because genetic code is nearly universal. Human insulin is used to treat diabetes.
Q6: Under which of the following conditions there will be no change in the reading frame of following mRNA? (NEET 2019)
5' AACAGCGGUGCUAUU 3'
(a) Deletion of GGU from 7th, 8th and 9th positions
(b) Insertion of G at 5th position
(c) Deletion of G from 5th position
(d) Insertion of A and G at 4th and 5th position respectively
Ans: (a)
In case of deletion of GGU from 7th, 8th and 9th position, there will be no change in reading frame of mRNA.

2018
Q1: Select the correct match. (NEET 2018)
(a) Alec Jeffreys - Streptococcus pneumoniae
(b) Alfred Hershey and Martha Chase - TMV
(c) Matthew Meselson and F. Stahl - Pisum sativum
(d) Francois Jacob and Jacques Monod - Lac operon
Ans: (d)
Francois Jacob and Jacque Monod proposed model of gene regulation known as operon model/lac operon. Alec Jeffreys gave DNA fingerprinting technique. Matthew Meselson and F. Stahl gave semiconservative DNA replication in E.coli. Alfred Hershey and Martha Chase proved DNA as genetic material not protein.
Q2: The experimental proof for semi-conservative replication of DNA was first shown in a (NEET 2018)
(a) Fungus
(b) Bacterium
(c) Plant
(d) Virus
Ans: (b)
Semi-conservative DNA replication was first shown in bacterium escherichia coli by Matthew Meselson and Franklin Stahl.
Q3: Select the correct statement. (NEET 2018)
(a) Franklin Stahl coined the term ‘‘ linkage ’’.
(b) Punnett square was developed by a British scientist.
(c) Spliceosomes take part in translation.
(d) Transduction was discovered by S. Altman.
Ans: (b)
- Punnet square was developed by a British geneticist Reginald C. Punnet.
- Franklin Stahl and Meselson proved semiconservative mode of DNA replication.
- Morgan coined the term linkage.
- Transduction was discovered by Zinder and Lederberg.
Q4: Select the correct match. (NEET 2018)
(a) Ribozyme - Nucleic acid
(b) F2 x Recessive parent - Dihybrid cross
(c) T.H. Morgan - Transduction
(d) G. Mendel - Transformation
Ans: (a)
Ribozyme is a catalytic RNA, which is nucleic acid.
Q5: Many ribosomes may associate with a single wiRNA to form multiple copies of a polypeptide simultaneously. Such strings of ribosomes are termed as (NEET 2018)
(a) Polysome
(b) Polyhedral bodies
(c) Plastidome
(d) Nucleosome
Ans: (a)
- Ribosomes may occur singly a monosomes or in rosettes and helical groups called polysomes. The different ribosomes of a polysome are connected with a strand of m-RNA.
- Nucleosome is a basic unit of DNA packaging in eukaryotes.
- Plastidome are the plastids of a cell when they are referred to a functional unit.
- Polyhedral bodies are involved in carbon fixation are present in autotrophic bacteria.
Q6: All of the following are part of an operon except (NEET 2018)
(a) An operator
(b) Structural genes
(c) An enhancer
(d) A promoter
Ans: (c)
Operon concept is for prokaryotes and enchancer sequences are present in eukaryotes.
Q7: AGGTATCGCAT is a sequence from the coding strand of a gene. What will be the corresponding sequence of the transcribed mRNA? (NEET 2018)
(a) AGGUAUCGCAU
(b) UGGTUTCGCAT
(c) ACCUAUGCGAU
(d) UCCAUAGCGUA
Ans: (a)
Coding strand and mRNA have same nucleotide sequence except, ‘T’ - Thymine is replaced by ‘U’ - Uracil in mRNA.
2017
Q1: The final proof for DNA as the genetic material came from the experiments of (NEET 2017)
(a) Hershey and Chase
(b) Avery, MacLeod and McCarty
(c) Hargobind Khorana
(d) Griffith
Ans: (a)
Hershey and Chase proved that DNA as genetic material. They used bacteriophage for their experiment.
Q2: If there are 999 bases in an RNA that code for a protein with 333 amino acids, and the base at position 901 is deleted such that the length of the RNA becomes 998 bases, how many codons will be altered? (NEET 2017)
(a) 11
(b) 33
(c) 333
(d) 1
Ans: (b)
If deletion happen at 901st position than the remaining 98 bases specifying for 33 codons of amino acids will be altered.
Q3: During DNA replication, Okazaki fragments are used to elongate (NEET 2017)
(a) The lagging strand towards replication fork
(b) The leading strand away from replication fork
(c) The lagging strand away from the replication fork
(d) The leading strand towards replication fork
Ans: (c)
- Two DNA polymerase molecules simultaneously work at the DNA fork, one on the leading strand and the other on the lagging strand.
- DNA polymerase synthesizes each Okazaki fragment at lagging strand in 5′-3′ direction. As the replication fork opens further, new Pkazaki fragments appear. The first Okazaki fragment appears away from the replication fork and thus the direction of elongation would be away from replication fork.
Q4: Which of the following RNAs should be most abundant in animal cell? (NEET 2017)
(a) tRNA
(b) mRNA
(c) miRNA
(d) rRNA
Ans: (d)
Ribosomal RNA (rRNA) is most abundant in animal cell. It constitutes 80% of total RNA of the cell.
Q5: Spliceosomes are not found in cells of (NEET 2017)
(a) Fungi
(b) Animals
(c) Bacteria
(d) Plants
Ans: (c)
Spliceosomes help in removal of introns. They will not occur in prokaryotes because prokaryotes do not have introns and thus, processing does not require splicing of mRNA.
Q6: The association of histone H1 with a nucleosome indicates that (NEET 2017)
(a) DNA replication is occurring
(b) The DNA is condensed into a chromatin fibre
(c) The DNA double helix is exposed
(d) Transcription is occurring
Ans: (b)
Histones help in packaging of DNA. In eukaryotes, DNA packaging is carried out with the help of positively charged basic proteins called histones. Histones are of five types - H1 H2A, H2B, H3 and H4. H1 is attached over the linker DNA. Histone contains a large proportion of the positively charged (basic) amino acids, lysine and arginine in their structure. DNA is negatively charged due to the phosphate groups on its backbone. The result of these opposite charges is strong attraction and therefore, high binding affinity between histones and DNA.
2016
Q1: Taylor conducted the experiments to prove semiconservative mode of chromosome replication on (NEET 2016 Phase 2)
(a) Vinca rosea
(b) Vicia faba
(c) Drosophila melanogaster
(d) E. coli
Ans: b
Taylor et al. (1957) conducted experiment on Vicia faba (broad bean) to prove semiconservative replication of DNA. He fed dividing cells of root tips of Vicia faba with radioactive 3H containing thymine instead of normal thymine and found that all the chromosomes became radioactive. Labelled thymine was then replaced with normal one. Next generation came to have radioactivity in one of the two chromatids of each chromosome while in subsequent generation radioactivity was present in 50% of the chromosomes. This is possible only if out of the two strands of a chromosome, one is formed afresh while the other is conserved at each replication.
Q2: The equivalent of a structural gene is (NEET 2016 Phase 2)
(a) Muton
(b) Cistron
(c) Operon
(d) Recon
Ans: (b)
Cislron (or gene) is a length ot DNA that contains the information for coding a specific Polypeptide chain or a functional RNA molecule transfer RNA or ribosomal RNA). Hence, cistron is a unit of function. Currently such a gene is called structural gene.
Q3: Which of the following rRNAs acts as structural RNA as well as ribozyme in bacteria? (NEET 2016 Phase 2)
(a) 5S rRNA
(b) 18S rRNA
(c) 23S rRNA
(d) 5.8S rRNA
Ans: (c)
23S rRNA acts as structural RNA as well as ribozyme in bacteria.
Q4: A molecule that can act as a genetic material must fulfill the traits given below, except (NEET 2016 Phase 2)
(a) It should be able to express itself in the form of ‘Mendelian characters’
(b) It should be able to generate its replica
(c) It should be unstable structurally and chemically
(d) It should provide the scope for slow changes that are required for evolution.
Ans: (c)
Genetic material should be structurally and chemically stable otherwise its expression will change and lead to loss of several metabolic functions, etc.
Q5: DNA-dependent RNA polymerase catalyses transcription on one strand of the DNA which is called the (NEET 2016 Phase 2)
(a) Template strand
(b) Coding strand
(c) Alpha strand
(d) Antistrand
Ans: (a)
The strand of DNA on which RNA polymerase binds to catalyse transcription is called template strand. It is also known as master or antisense strand. It lias the polarity of 3' → 5'.
Q6: Which one of the following is the starter codon? (NEET 2016 Phase 1)
(a) UAA
(b) UAG
(c) AUG
(d) UGA
Ans: (c)
The start codon is the first codon of a messenger RNA (mRNA) transcript translated by a ribosome. The start codon always codes for methionine in eukaryotes and a modified Met (fMet) in prokaryotes. The most common start codon is AUG.
Q7: Which of the following is required as inducer (s) for the expression of Lac operon? (NEET 2016 Phase 1)
(a) Lactose
(b) Lactose and Galactose
(c) Glucose
(d) Galactose
Ans: (a)
In Lac operon, lactose is an inducer. It binds with suppressor and inactivates it. It allows RNA polymers access to the promoter and transcription proceeds.
Q8: A complex of ribosomes attached to a single strand of RNA is known as (NEET 2016 Phase 1)
(a) Polypeptide
(b) Okazaki fragment
(c) Polysome
(d) Polymer
Ans: (c)
A polysome or polyribosome is a complex of an mRNA molecule and two or more ribosomes, which is formed during the active translation process. They were initially named as ergosomes in 1963. However, further research by Jonathan Warner and Alex Rich characterized polysome
|
78 videos|276 docs|174 tests
|
FAQs on NEET Previous Year Questions(2016-24): Molecular Basis of Inheritance - Biology Class 12
| 1. What is the role of DNA replication in the molecular basis of inheritance? |  |
| 2. How do mutations contribute to genetic variation in organisms? |  |
| 3. What are the different types of gene mutations that can occur in DNA? |  |
| 4. How does DNA packaging and epigenetic modifications regulate gene expression? |  |
| 5. How is the central dogma of molecular biology related to the process of gene expression? |  |

|
Explore Courses for NEET exam
|

|


















