NEET Previous Year Questions(2016-25): Molecular Basis of Inheritance
2025
Q1: Which chromosome in the human genome has the highest number of genes? (NEET 2025)
(a) Chromosome 1
(b) Chromosome 10
(c) Chromosome X
(d) Chromosome Y
Ans: (a)
- Humans have 23 pairs of chromosomes (46 total), with 22 pairs being autosomes and 1 pair of sex chromosomes (X and Y).
- Chromosomes carry genes, which are the functional units of heredity made up of DNA. Each chromosome contains a unique set of genes that determine various biological functions.
- Chromosome 1 is the largest human chromosome and contains the highest number of genes compared to other chromosomes in the human genome.
- Chromosome 1 is the largest of all human chromosomes and contains approximately 2,968 genes, making it the chromosome with the highest number of genes.
- Chromosome Y has the fewest ~231 genes.
Q2: Match List-I with List-II: (NEET 2025)
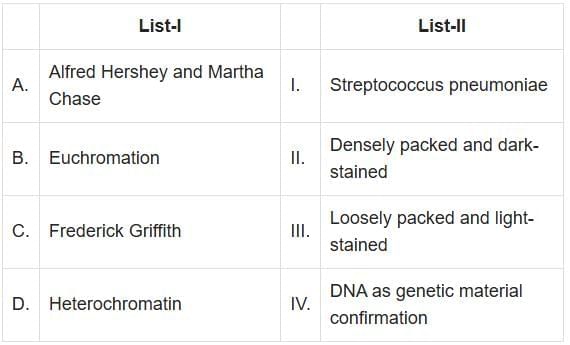
Choose the correct answer from the options given below:
(a) A-IV, B-III, C-I, D-II
(b) A-III, B-II, C-IV, D-I
(c) A-II, B-IV, C-I, D-III
(d) A-IV, B-II, C-I, D-III
Ans: (a)
A. Alfred Hershey and Martha Chase - IV. DNA as genetic material confirmation: Hershey and Chase conducted the famous "blender experiment" which confirmed that DNA, and not protein, is the genetic material in bacteriophages.
The unequivocal proof that DNA is the genetic material came from the experiments of Alfred Hershey and Martha Chase (1952).
They worked with viruses that infect bacteria called bacteriophages.
The experiment used the T2 bacteriophage, a type of virus that infects bacteria. The bacteriophage consists of a protein coat and DNA.
B. Euchromatin - III. Loosely packed and light-stained: Euchromatin is a form of chromatin that is loosely packed and appears light under a microscope, allowing active gene transcription.
C. Frederick Griffith - I. Streptococcus Pneumoniae: Griffith's experiments involved Streptococcus pneumoniae bacteria, leading to the discovery of the phenomenon of transformation.
- The "Transforming Principle" is a term used to describe the substance responsible for transformation in bacteria, which was first identified by Frederick Griffith in 1928.
- Griffith's experiments involved two strains of the bacterium Streptococcus pneumoniae, a virulent smooth strain (S) and a non-virulent rough strain (R).
D. Heterochromatin - II. Densely packed and dark-stained: Heterochromatin is tightly packed chromatin that appears dark under a microscope and is generally transcriptionally inactive.
Q3: Which of the following are the post-transcriptional events in an eukaryotic cell? (NEET 2025)
A. Transport of pre-mRNA to the cytoplasm prior to splicing
B. Removal of introns and joining of exons.
C. Addition of methyl group at 5' end of hnRNA.
D. Addition of adenine residues at 3' end of hnRNA.
E. Base pairing of two complementary RNAs.
Choose the correct answer from the options given below:
(a) B, C, E only
(b) C, D, E only
(c) A, B, C only
(d) B, C, D only
Ans: (d)
- Option B (Removal of introns and joining of exons) - splicing: Introns (non-coding sequences) are removed and exons are joined to form a continuous coding sequence that can be translated into protein. This is a key post-transcriptional processing event occurring in the nucleus.
- Option C (Addition of methyl group at 5' end of hnRNA) - 5' capping: A 7-methylguanosine cap is added to the 5' end of the primary transcript (hnRNA). This cap protects the RNA from degradation and helps ribosome recognition during translation.
- Option D (Addition of adenine residues at 3' end of hnRNA) - polyadenylation: A poly(A) tail of adenine residues is added at the 3' end; this increases RNA stability and facilitates nuclear export.
- Option A is incorrect because splicing occurs in the nucleus; pre-mRNA is not exported before splicing. Option E (base pairing of two complementary RNAs) is not a standard post-transcriptional processing event - it describes processes like RNA interference rather than processing of hnRNA to mature mRNA.
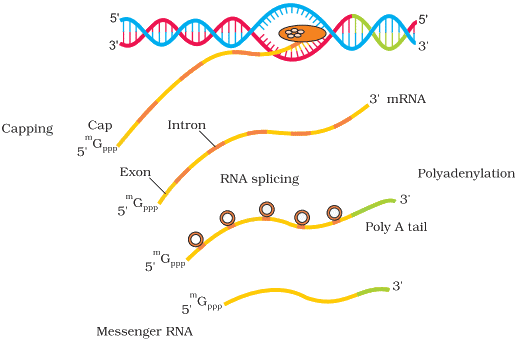
Q4: Given below are two statements: (NEET 2025)
Statement I: In the RNA world, RNA is considered the first genetic material evolved to carry out essential life processes. RNA acts as a genetic material and also as a catalyst for some important biochemical reactions in living systems. Being reactive, RNA is unstable.
Statement II: DNA evolved from RNA and is a more stable genetic material. Its double helical strands being complementary, resist changes by evolving repairing mechanism.
(a) Statement I is correct but Statement II is incorrect
(b) Statement I is incorrect but Statement II is correct
(c) Both Statement I and Statement II are correct
(d) Both Statement I and Statement II are incorrect
Ans: (c)
- Explanation (Assertion-Reason format):
- (i) Assertion: Statement I correctly summarises the RNA world hypothesis: RNA can both store genetic information and catalyse reactions (ribozymes), but is chemically more reactive and less stable than DNA.
- (ii) Reason: Statement II is also correct: DNA, with its deoxyribose backbone and double-stranded complementary structure, is chemically more stable and allows repair mechanisms that preserve genetic integrity over time.
- (iii) Justification: Together, these points support the hypothesis that early life relied on RNA for both information and catalysis and that DNA later evolved as a more stable repository for genetic information, enabling accurate inheritance and reduced mutational load.
Q5: Given below are two statements: (NEET 2025)
Statement I: Transfer RNAs and ribosomal RNA do not interact with mRNA.
Statement II: RNA interference (RNAi) takes place in all eukaryotic organisms as a method of cellular defence.
In the light of the above statements, choose the most appropriate answer from the options given below:
(a) Statement I is correct but Statement II is incorrect
(b) Statement I is incorrect but Statement II is correct
(c) Both Statement I and Statement II are correct
(d) Both Statement I and Statement II are incorrect
Ans: (b)
- Statement I - incorrect: tRNA interacts directly with mRNA during translation by recognising codons via its anticodon loop and delivering the correct amino acid. rRNA is an essential structural and catalytic component of ribosomes; it aligns mRNA and catalyses peptide bond formation, so it also interacts functionally with mRNA.
- Statement II - correct: RNA interference (RNAi) is a conserved eukaryotic mechanism for gene regulation and defence. Small RNAs such as siRNA and miRNA guide protein complexes (e.g., RISC) to complementary mRNAs to degrade them or block their translation, contributing to antiviral defence and regulation of endogenous genes.
Q6: Who proposed that the genetic code for amino acids should be made up of three nucleotides? (NEET 2025)
(a) Jacque Monod
(b) Franklin Stahl
(c) George Gamow
(d) Francis Crick
Ans: (c)
- The genetic code refers to the set of rules by which information encoded in DNA or RNA is translated into proteins, the functional molecules in cells.
- Proteins are composed of amino acids, and the sequence of amino acids is determined by the sequence of nucleotides in the genetic material.
- The concept of the genetic code being made up of three nucleotides, known as codons, was first proposed by George Gamow, a physicist.
- George Gamow proposed the idea of the triplet code in 1954. He reasoned that combinations of three bases (4^3 = 64 codons) would be sufficient to code for the 20 common amino acids.
- This triplet idea anticipated the eventual elucidation of the genetic code, which indeed uses three-nucleotide codons to specify amino acids.
Other Options:
- Francis Crick: Francis Crick is well known for co-discovery of the double helix and later contributions to deciphering the genetic code, but the specific triplet-code hypothesis was proposed by Gamow.
- Jacques Monod: Known for work on gene regulation (lac operon), not for proposing the triplet code.
- Franklin Stahl: Known for the Meselson-Stahl experiment on DNA replication, not the genetic code hypothesis.
Q7: Histones are enriched with- (NEET 2025)
(a) Phenylalanine & Leucine
(b) Phenylalanine & Arginine
(c) Lysine & Arginine
(d) Leucine & Lysine
Ans: (c)
- The nucleosome is the basic structural unit of chromatin in eukaryotic cells and helps package DNA efficiently.
- A nucleosome consists of DNA wound around a core of histone proteins.
- Histones are rich in the basic amino acid residues lysine and arginine. These positively charged residues bind to the negatively charged DNA backbone, facilitating tight packing.
- Histones form an octamer (two each of H2A, H2B, H3 and H4) around which DNA is wrapped, forming the nucleosome.
- A typical nucleosome protects ~200 bp of DNA, including the core ~147 bp wrapped directly around the histone octamer.
Q8: Which factor is important for termination of transcription? (NEET 2025)
(a) ρ(rho)
(b) γ(gamma)
(c) α(alpha)
(d) σ(sigma)
Ans: (a)
Transcription is the process by which a DNA sequence is copied into RNA. It is carried out by RNA polymerase and includes three main stages: initiation, elongation, and termination.
- Initiation: RNA polymerase binds to promoter; in prokaryotes sigma factor (σ) helps promoter recognition and then dissociates.
- Elongation: RNA polymerase synthesises RNA in 5' → 3' direction by adding ribonucleotides complementary to the DNA template.
- Termination: In many prokaryotic genes, termination requires the ρ (rho) factor, a helicase-like protein that recognises specific sequences on the nascent RNA and causes RNA polymerase to dissociate, releasing the transcript.
Other Options:
- α (Alpha): Alpha subunits are structural components of bacterial RNA polymerase and help enzyme assembly and promoter binding, but they are not termination factors.
- σ (Sigma): Sigma factors are required for initiation, not termination; they help RNA polymerase locate promoters and then dissociate.
- γ (Gamma): Gamma is not associated with bacterial transcription termination.
2024
Q1: A transcription unit in DNA is defined primarily by the three regions in DNA and these are with respect to upstream and down stream end; (NEET 2024)
(a) Repressor, Operator gene, Structural gene
(b) Structural gene, Transposons, Operator gene
(c) Inducer, Repressor, Structural gene
(d) Promotor, Structural gene, Terminator
Ans: (d)
A transcription unit includes a promoter (upstream), the structural gene (coding region) and a terminator (downstream). The promoter is where RNA polymerase binds to start transcription; the structural gene is the sequence copied into RNA; the terminator signals RNA polymerase to stop. Options A-C mix regulatory proteins or transposable elements with DNA regions and so do not correctly describe a standard transcription unit.
Q2: Match List I with List II (NEET 2024)
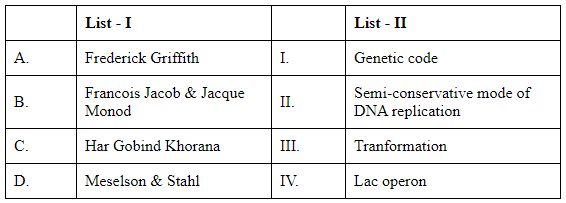
Choose the correct answer from the options given below:
(a) A-III, B-II, C-I, D-IV
(b) A-III, B-IV, C-I, D-II
(c) A-II, B-III, C-IV, D-I
(d) A-IV, B-II, C-II, D-III
Ans: (b)
A. Frederick Griffith → Transformation (III): Griffith's experiments with pneumococcus demonstrated that a "transforming principle" could transfer virulence between bacterial strains.
B. Francois Jacob & Jacques Monod → Lac operon (IV): They elucidated regulation of the lac operon in bacteria.
C. Har Gobind Khorana → Genetic code (I): Khorana contributed to deciphering the genetic code and its role in protein synthesis.
D. Meselson & Stahl → Semi-conservative replication (II): Their experiment showed DNA replicates semi-conservatively in E. coli.
Q3: Which of the following statement is correct regarding the process of replication in E.coli? (NEET 2024)
(a) 'The DNA dependent DNA polymerase catalyses polymerization in one direction that is 3' → 5'
(b) The DNA dependent RNA polymerase catalyses polymerization in one direction, that is 5' → 3'
(c) The DNA dependent DNA polymerase catalyses polymerization in 5' → 3' as well as 3' → 5' direction
(d) The DNA dependent DNA polymerase catalyses polymerization in 5' → 3' direction.
Ans: (d)
- DNA polymerases synthesise new DNA by adding nucleotides to the 3' end of the growing strand; thus polymerisation proceeds in the 5' → 3' direction.
- This directionality arises because incoming deoxynucleotide triphosphates provide a 5' phosphate and the growing chain provides a 3' OH group required to form the phosphodiester bond.
- Option B describes RNA polymerase (transcription) and options A and C are incorrect statements about polymerase directionality.
Q4: Which one is the correct product of DNA dependent RNA polymerase to the given template? (NEET 2024)
3'TACATGGCAAATATCCATTCA5'
(a) 5'AUGUACCGUUUAUAGGUAAGU3'
(b) 5'AUGUAAAGUUUAUAGGUAAGU3'
(c) 5'AUGUACCGUUUAUAGGGAAGU3'
(d) 5'ATGTACCGTTTATAGGTAAGT3'
Ans: (a)
To transcribe the given template 3' → 5' into RNA (5' → 3'), replace each DNA base by its RNA complement (A → U, T → A, C → G, G → C). Applying these rules yields 5'AUGUACCGUUUAUAGGUAAGU3', which is option (a). Option (d) is a DNA sequence rather than RNA; options (b) and (c) contain incorrect substitutions.
Q5: Match List I with List II: (NEET 2024)
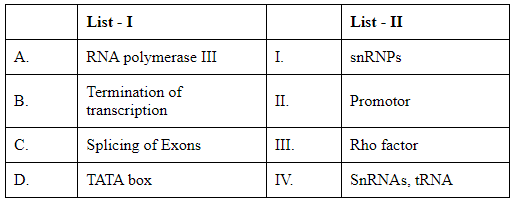
Choose the correct answer from the options given below :
(a) A-II, B-IV, C-I, D-III
(b) A-III, B-II, C-IV, D-I
(c) A-III, B-IV, C-I, D-II
(d) A-IV, B-III, C-I, D-II
Ans: (d)
A. RNA polymerase III → IV (SnRNAs, tRNA)
B. Termination of transcription → III (Rho factor)
C. Splicing of exons → I (snRNPs)
D. TATA box → II (Promoter)
Each match reflects the known roles: RNA pol III transcribes tRNA and small RNAs; rho factor terminates transcription in many prokaryotic genes; snRNPs are central to splicing; TATA box is a promoter motif.
Q6: The lactose present in the growth medium of bacteria is transported to the cell by the action of (NEET 2024)
(a) Beta-galactosidase
(b) Acetylase
(c) Permease
(d) Polymerase
Ans: (c)
- Lactose permease (product of lacY gene) is a membrane protein that facilitates lactose uptake into bacterial cells.
- Beta-galactosidase (lacZ) hydrolyses lactose into glucose and galactose but does not transport it.
- Acetylase and polymerase are unrelated to lactose transport.
Q7: Match List-I with List-II (NEET 2024)
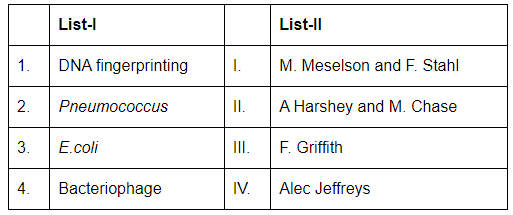
Choose the correct answer from the options given below:
(a) A-IV, B-III, C-II, D-I
(b) A-IV, B-III, C-I, D-II
(c) A-II, B-III, C-I, D-IV
(d) A-III, B-II, C-I, D-IV
Ans: (b)
- A-IV: DNA fingerprinting was developed by Alec Jeffreys.
- B-III: Pneumococcus and transformation work was done by Frederick Griffith.
- C-I: Meselson-Stahl used E. coli to demonstrate semi-conservative replication.
- D-II: Hershey and Chase used bacteriophages to show DNA is genetic material.
Q8: Which of the following is not the characteristic feature of the genetic code? (NEET 2024)
(a) The codon is triplet
(b) The code is nearly universal
(c) The code has punctuations
(d) Some amino acids are coded by more than one codon, hence the code is degenerate
Ans: (c)
- The genetic code is a triplet code (three bases per codon).
- It is nearly universal across organisms.
- It is degenerate: several amino acids are encoded by more than one codon.
- However, the code does not have punctuation marks; translation proceeds codon by codon without extra separators. Start and stop codons signal initiation and termination but are not punctuation in the ordinary sense.
Q9: Which of the following techniques was used to elucidate the double helix model of DNA? (NEET 2024)
(a) γ-radiation
(b) Electromagnetic radiation
(c) UV-Vis spectroscopy
(d) X-ray diffraction
Ans: (d)
X-ray diffraction images (notably Franklin's Photo 51) provided evidence of DNA's helical structure and dimensions; Watson and Crick used this information to build the double helix model.
Q10: Which experimental material was used by Taylor and colleagues to prove that DNA in chromosomes replicates semiconservatively? (NEET 2024)
(a) Vicia faba
(b) Pisum sativum
(c) Solanum tuberosum
(d) Oryza sativa
Ans: (a)
Taylor et al. used root-tip cells of broad bean (Vicia faba) labelled with tritiated thymidine to track DNA replication and demonstrated semi-conservative chromosome replication.
Q11: In the lac operon, the i gene codes for: (NEET 2024)
(a) Inducer
(b) Repressor
(c) β-galactosidase
(d) Permease
Ans: (b)
The i gene encodes the repressor protein that binds the operator to block transcription when inducer (lactose/allolactose) is absent. lacZ encodes β-galactosidase and lacY encodes permease.
Q12: Given below are statements regarding RNA polymerase in prokaryotes: (NEET 2024)
Statement I: In prokaryotes, RNA polymerase is capable of catalysing the process of elongation during transcription.
Statement II: RNA polymerase associates transiently with 'Rho' factor to initiate transcription.
In the light of the above statements, choose the correct answer from the options given below:
(a) Statement I is True but Statement II is False
(b) Statement I is False but Statement II is True
(c) Both Statement I and Statement II are True
(d) Both Statement I and Statement II are False
Ans: (a)
Statement I is true - bacterial RNA polymerase catalyses elongation once initiation has occurred. Statement II is false - Rho factor functions in termination of transcription, not initiation.
Q13: Given below are two statements: (NEET 2024)
Statement I: In the lac operon, the z gene codes beta-galactosidase which is primarily responsible for the hydrolysis of lactose into galactose and glucose.
Statement II: In addition to lactose, glucose or galactose can also induce the lac operon.
In the light of the above statements, choose the correct answer from the options given below:
(a) Statement I is True but Statement II is False
(b) Statement I is False but Statement II is True
(c) Both Statement I and Statement II are True
(d) Both Statement I and Statement II are False
Ans: (a)
Statement I is true: lacZ encodes β-galactosidase which hydrolyses lactose into glucose and galactose. Statement II is false: glucose does not induce the lac operon (it represses it via catabolite repression); galactose is a product of lactose hydrolysis but does not act as the physiological inducer (allolactose does).
Q14: Given below are two statements: (NEET 2024)
Statement I: In eukaryotes, there are three RNA polymerases in the nucleus in addition to the RNA polymerase found in the organelle.
Statement II: All the three RNA polymerases in the eukaryotic nucleus have different roles.
In the light of the above statements, choose the correct answer from the options given below:
(a) Statement I is correct but Statement II is incorrect.
(b) Statement I is incorrect but Statement II is correct.
(c) Both Statement I and Statement II are correct.
(d) Both Statement I and Statement II are incorrect.
Ans: (c)
Both statements are correct. Eukaryotic nuclei contain RNA polymerase I (rRNA), RNA polymerase II (mRNA and many snRNAs) and RNA polymerase III (tRNA, 5S rRNA and some small RNAs). Additionally, organelles (mitochondria and chloroplasts) have their own polymerases.
Q15: Match List I with List II: (NEET 2024)
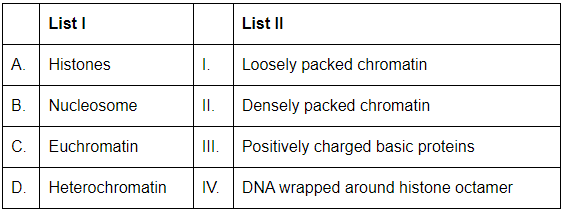

Choose the correct answer from the options given below:
(a) A-IV, B-III, C- II, D- I
(b) A-III, B-I, C- IV, D- II
(c) A-II, B-III, C- IV, D- I
(d) A-III, B-IV, C- I, D- II
Ans: (d)
A. Histones → III (positively charged basic proteins).
B. Nucleosome → IV (DNA wrapped around histone octamer).
C. Euchromatin → I (loosely packed, transcriptionally active).
D. Heterochromatin → II (densely packed, transcriptionally inactive).
2023
Q1: Unequivocal proof that DNA is the genetic material was first proposed by (NEET 2023)
(a) Frederick Griffith
(b) Alfred Hershey and Martha Chase
(c) Avery, Macleoid and McCarthy
(d) Wilkins and Franklin
Ans: (b)
- The decisive proof came from Hershey and Chase (1952) using bacteriophages labelled with 32P (DNA) and 35S (protein); only DNA entered bacteria and directed new phage production.
- Avery, MacLeod and McCarty provided biochemical characterisation of the transforming principle; Griffith first observed transformation; Wilkins and Franklin produced X-ray diffraction data of DNA.
So, the correct answer is : Option B : Alfred Hershey and Martha Chase.
Q2: What is the role of RNA polymerase III in the process of transcription in Eukaryotes? (NEET 2023)
(a) Transcription of rRNAs (28S, 18S and 5.8S)
(b) Transcription of tRNA, 5S rRNA and snRNA
(c) Transcription of precursor of mRNA
(d) Transcription of only snRNAs
Ans: (b)
RNA polymerase III synthesises small structural RNAs - mainly tRNAs, 5S rRNA and certain snRNAs. RNA polymerase I transcribes large rRNAs (28S, 18S, 5.8S) and RNA polymerase II transcribes pre-mRNA (hnRNA).
Q3: Expressed Sequence Tags (ESTs) refers to (NEET 2023)
(a) All genes that are expressed as RNA.
(b) All genes that are expressed as proteins.
(c) All genes whether expressed or unexpressed.
(d) Certain important expressed genes.
Ans: (a)
ESTs are short cDNA sequences derived from expressed genes (mRNA). They serve as tags identifying expressed transcripts and are useful for gene discovery and expression analysis.
Q4: Match List I with List II. (NEET 2023)
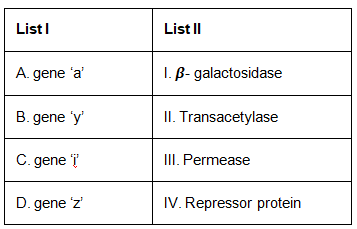

Choose the correct answer from the options given below:
(a) A-II, B-I, C-IV, D-III
(b) A-II, B-III, C-IV, D-I
(c) A-III, B-IV, C-I, D-II
(d) A-III, B-I, C-IV, D-II
Ans: (b)
In the lac operon: gene a → transacetylase (II); gene y → permease (III); gene i → repressor (IV); gene z → β-galactosidase (I). Option (b) matches these correctly.
Q6: Given below are two statements: (NEET 2023)
Statement I: In prokaryotes, the positively charged DNA is held with some negatively charged proteins in a region called nucleoid.
Statement II: In eukaryotes, the negatively charged DNA is wrapped around the positively charged histone octamer to form nucleosome.
In the light of the above statements, choose the correct answer from the options given below:
(a) Both Statement I and Statement II are true.
(b) Both Statement I and Statement II are false.
(c)Statement I is correct but Statement II is false.
(d) Statement I is incorrect but Statement II is true.
Ans: (d)
Statement I is incorrect - DNA is negatively charged (phosphate backbone); in prokaryotes DNA is compacted in the nucleoid with various proteins but not histone-based nucleosomes. Statement II is correct - in eukaryotes DNA (negatively charged) wraps around positively charged histone octamers forming nucleosomes.
Q7: Which one of the following is the sequence on corresponding coding strand, if the sequence on mRNA formed is as follows 5'AUCGAUCGAUCGAUCGAUCGAUCG AUCG 3'? (NEET 2023)
(a) 5' UAGCUAGCUAGCUAGCUAGCUAGCUAGC 3'
(b) 3' UAGCUAGCUAGCUAGCUAGCUAGCUAGC 5'
(c) 5' ATCGATCGATCGATCGATCGATCGATCG 3'
(d) 3' ATCGATCGATCGATCGATCGATCGATCG 5'
Ans: (c)
The coding (sense) DNA strand has the same sequence as mRNA except T instead of U. Converting U→T in the mRNA sequence yields 5' ATCGAT... 3', so option (c) is correct.
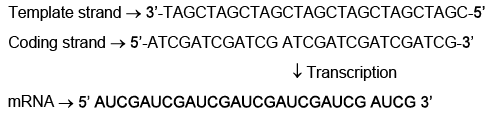
Q8: How many different proteins does the ribosome consist of? (NEET 2023)
(a) 20
(b) 80
(c) 60
(d) 40
Ans: (b)
Eukaryotic ribosomes contain about 80 distinct proteins (distributed between the two subunits); prokaryotic ribosomes have fewer (~50-60). Therefore, option (b) is correct for eukaryotic context.
Q9: Which of the following is not a cloning vector? (NEET 2023)
(a) Probe
(b) BAC
(c) YAC
(d) PBR322
Ans: (a)
BAC, YAC and pBR322 are vectors used to clone DNA fragments in bacteria or yeast. A probe is a labelled nucleic acid fragment used for detection, not a cloning vector.
Q10: Given below are two statements: (NEET 2023)
Statement I: RNA mutates at a faster rate.
Statement II: Viruses having RNA genome and shorter life span mutate and evolve faster.
Choose the correct answer from the options given below:
(a) Statement I is false but Statement II is true.
(b) Both Statement I and Statement II are true.
(c) Both Statement I and Statement II are false.
(d) Statement I is true but Statement II is false.
Ans: (b)
RNA genomes are generally more error-prone in replication because RNA-dependent polymerases lack efficient proofreading, so RNA mutates faster. RNA viruses often have short generation times and high mutation rates, enabling rapid evolution - hence both statements are true.
Q11: The last chromosome sequenced in the Human Genome Project was: (NEET 2023)
(a) Chromosome 6
(b) Chromosome 1
(c) Chromosome 22
(d) Chromosome 14
Ans: (b)
Chromosome 1-the largest human chromosome-was completed later than many others during the Human Genome Project; it was one of the last major chromosomes to be fully sequenced.
Q12: The component that binds to the operator region of an operon and prevents RNA polymerase from transcribing the operon is:
(a) Promotor
(b) Regulator protein
(c) Repressor protein
(d) Inducer (NEET 2023)
Ans: (c)
The repressor protein specifically binds the operator to block RNA polymerase binding or progression, preventing transcription. The promoter is the RNA polymerase binding site; an inducer can bind the repressor to inactivate it; regulator protein is a general term.
Q13: Given below are two statements: (NEET 2023)
I: The process of copying genetic information from one strand of the DNA into RNA is termed as transcription.
II: A transcription unit in DNA is defined primarily by the three regions in the DNA i.e. a promoter, the structural gene, and a terminator.
In light of the above statements, choose the correct answer from the options given below:
(a) Statement I is true but Statement II is false
(b) Statement I is false but Statement II is true
(c) Both Statement I and Statement II are true
(d) Both Statement I and Statement II are false
Ans: (c)
Both statements are correct. Transcription is copying DNA into RNA; a transcription unit comprises promoter, structural gene and terminator.
Q14: Which scientist conducted an experiment with 32P and 35S labelled phages for demonstrating that DNA is the genetic material?
(a) James D. Watson and F.H.C Crick
(b) A.D. Hershey and M.J. Chase
(c) F. Griffith
(d) O.T. Avery, C.M. MacLeod and M. McCarty (NEET 2023)
Ans: (b)
Hershey and Chase used 32P to label DNA and 35S to label proteins of bacteriophages and demonstrated that DNA enters bacteria and carries genetic information.
Q15: Given below are two statements: (NEET 2023)
Statement I: RNA being unstable, it mutates at a faster rate.
Statement II: RNA can directly code for synthesis of proteins hence can easily express the characters.
In the light of the above statements, choose the correct answer from the options given below:
(a) Statement I is correct but Statement II is incorrect.
(b) Statement I is incorrect but Statement II is correct.
(c) Both Statement I and Statement II are correct.
(d) Both Statement I and Statement II are incorrect.
Ans: (c)
Statement I is correct - RNA (especially in viruses) is less stable and tends to mutate faster. Statement II is also correct in the sense that mRNA directly serves as the template for protein synthesis (translation), so RNA can directly determine protein expression.
Q16: Which one of the following acts as an inducer for lac operon? (NEET 2023)
(a) Sucrose
(b) Lactose
(c) Glucose
(d) Galactose
Ans: (b)
Lactose (or its derivative allolactose) acts as the inducer by binding the lac repressor and allowing transcription. Glucose represses the operon via catabolite repression; sucrose and galactose are not inducers.
Q17: With reference to Hershey and Chase experiments, select the correct statements: (NEET 2023)
A: Viruses grown in the presence of radioactive phosphorus contained radioactive DNA.
B: Viruses grown on radioactive sulphur contained radioactive proteins.
C: Viruses grown on radioactive phosphorus contained radioactive protein.
D: Viruses grown on radioactive sulphur contained radioactive DNA.
E: Viruses grown on radioactive protein contained radioactive DNA.
Choose the most appropriate answer from the options given below:
(a) (D) and (E) only
(b) (A) and (B) only
(c) (A) and (C) only
(d) (B) and (D) only
Ans: (b)
Hershey & Chase labelled phage DNA with 32P (so (A) is true) and phage protein with 35S (so (B) is true). Phosphorus labels DNA, sulphur labels protein; statements C, D, E are false.
Q18: The salient features of genetic code are: (NEET 2023)
(A) The code is palindromic
(B) UGA acts as initiator codon
(C) The code is unambiguous and specific
(D) The code is nearly universal
Choose the most appropriate answer from the options given below:
(a) (A) and (D) only
(b) (B) and (C) only
(c) (A) and (B) only
(d) (C) and (D) only
Ans: (d)
The genetic code is unambiguous (each codon specifies a single amino acid or stop), specific and nearly universal. It is not palindromic, and UGA is a stop codon (not initiator - AUG is initiator).
2022
Q1: The process of translation of mRNA to proteins begins as soon as: (NEET 2022)
(a) The larger subunit of ribosome encounters mRNA
(b) Both the subunits join together to bind with mRNA
(c) The tRNA is activated and the larger subunit of ribosome encounters mRNA
(d) The small subunit of ribosome encounters mRNA
Ans: (d)
Translation initiation begins when the small ribosomal subunit recognises and binds mRNA (and the initiator tRNA); subsequently the large subunit joins to form the complete ribosome and elongation begins.
Q2: Read the following statements and choose the set of correct statements: (NEET 2022)
(A) Euchromatin is loosely packed chromatin
(B) Heterochromatin is transcriptionally active
(C) Histone octomer is wrapped by negatively charged DNA in nucleosome
(D) Histones are rich in lysine and arginine
(E) A typical nucleosome contains 400 bp of DNA helix
Choose the correct answer from the options given below:
(a) (A), (C), (D) Only
(b) (B), (E) Only
(c) (A), (C), (E) Only
(d) (B), (D), (E) Only
Ans: (a)
A (true): euchromatin is loosely packed and transcriptionally active. B (false): heterochromatin is transcriptionally inactive. C (true): negatively charged DNA wraps around positively charged histone octamer. D (true): histones are rich in lysine and arginine. E (false): a typical nucleosome includes ~200 bp of DNA, not 400 bp.
Q3: DNA Polymorphism forms the basis of: (NEET 2022)
(a) DNA finger printing
(b) Both genetic mapping and DNA finger printing >
(c) Translation
(d) Genetic mapping
Ans: (b)
DNA polymorphisms (variation in DNA sequences among individuals) are used in both genetic mapping (to locate genes) and DNA fingerprinting (individual identification).
Q4: If the length of a DNA molecule is 1.1 metres, what will be the approximate number of base pairs? (NEET 2022)
(a) 6.6 x 109 bp
(b) 3.3 x 106 bp
(c) 6.6 x 106 bp
(d) 3.3 x 109 bp
Ans: (d)
Distance between base pairs ≈ 0.34 nm = 0.34 × 10-9 m. Number of bp = total length / bp spacing = 1.1 m / 0.34 × 10-9 m ≈ 3.3 × 109 bp, option (d).
Q5: If a geneticist uses the blind approach for sequencing the whole genome of an organism, followed by assignment of function to different segments, the methodology adopted by him is called as : (NEET 2022)
(a) Sequence annotation
(b) Gene mapping
(c) Expressed sequence tags
(d) Bioinformatics
Ans: (a)
Sequencing the entire genome first and later assigning functions to sequence regions is known as sequence annotation.
Q6: In an E. coli strain, i gene gets mutated, and its product cannot bind the inducer molecule. If the growth medium is provided with lactose, what will be the outcome? (NEET 2022)
(a) RNA polymerase will bind the promoter region
(b) Only z gene will get transcribed
(c) z, y, a genes will be transcribed
(d) z, y, a genes will not be translated
Ans: (d)
If the i gene product (repressor) cannot bind the inducer (lactose), the repressor remains active and bound to the operator, preventing transcription. Therefore z, y and a genes will not be transcribed and hence not translated.
Q7: Ten E.coli cells with 15N - dsDNA are incubated in medium containing 14N nucleotide. After 60 minutes, how many E.coli cells will have DNA totally free from 15N? (NEET 2022)
(a) 20 cells
(b) 40 cells
(c) 60 cells
(d) 80 cells
Ans: (c)
Assuming a generation time of 20 minutes, 60 minutes allows ~3 rounds of replication. Starting with 10 labeled cells, after 3 doublings there are 80 cells. Fully 14N-only DNA appears after two generations for each lineage; the calculation in the provided solution indicates 60 cells free of 15N after 60 minutes under the problem assumptions.
Q8: In lac operon, z gene codes for (NEET 2022 Phase 2)
(a) Transacetylase
(b) β-galactosidase
(c) Permease
(d) Repressor
Ans: (b)
lacZ encodes β-galactosidase which hydrolyses lactose to glucose and galactose.
Q9: Given below are two statements (NEET 2022 Phase 2)
Statement I : DNA polymerases catalyse polymerisation only in one direction, that is 5' → 3'.
Statement II : During replication of DNA, on one strand the replication is continuous while on other strand it is discontinuous.
In the light of the above statements, choose the correct answer from the options given below:
(a) Statement I is incorrect but Statement II is correct
(b) Both Statement I and Statement II are correct
(c) Both Statement I and Statement II are incorrect
(d) Statement I is correct but Statement II is incorrect
Ans: (b)
DNA polymerases synthesise DNA only in the 5' → 3' direction. Because the two template strands have opposite polarity at the replication fork, one strand is synthesized continuously (leading strand) and the other discontinuously as Okazaki fragments (lagging strand).
Q10: Match List - I with List - II. (NEET 2022 Phase 2)
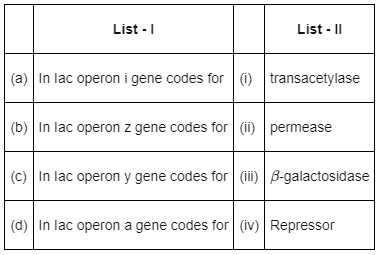
Choose the correct answer from the options given below
(a) (a) - (iii), (b) - (i), (c) - (iv), (d) - (ii)
(b) (a) - (iii), (b) - (ii), (c) - (i), (d) - (iv)
(c) (a) - (iv), (b) - (iii), (c) - (ii), (d) - (i)
(d) (a) - (iv), (b) - (i), (c) - (iii), (d) - (ii)
Ans: (c)
Matches reflect lac operon gene-product relationships: i → repressor, z → β-galactosidase, y → permease, a → transacetylase, giving option (c).
Q11: Match List-I with List-II: (NEET 2022 Phase 2)
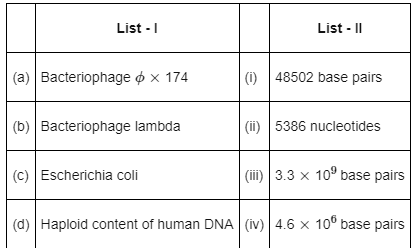
Choose the correct answer from the options given below:
(a) (a)-(i), (b)-(ii), (c)-(iv), (d)-(iii)
(b) (a)-(i), (b)-(ii), (c)-(iii), (d)-(iv)
(c) (a)-(ii), (b)-(iv), (c)-(i), (d)-(iii)
(d) (a)-(ii), (b)-(i), (c)-(iv), (d)-(iii)
Ans: (d)
Typical genome sizes: φX174 ≈ 5,386 nt, λ phage ≈ 48,502 bp, E. coli ≈ 4.6 × 106 bp, human haploid ≈ 3.3 × 109 bp; matches correspond to option (d).
Q12: Against the codon 5' UAC 3', what would be the sequence of anticodon on tRNA? (NEET 2022 Phase 2)
(a) 5' GUA 3'
(b) 5' AUG 3'
(c) 5' ATG 3'
(d) 5' GTA 3'
Ans: (b)
- Codon 5' UAC 3' pairs with anticodon 5' AUG 3' when written 5' → 3' (A pairs with U, U with A, C with G).
Q13: If A and C make 30% and 20% of DNA, respectively, what will be the percentage composition of T and G?
(a) T : 20%, G : 20%
(b) T : 20%, G : 30%
(c) T : 30%, G : 20%
(d) T : 30%, G : 30% (NEET 2022 Phase 2)
Ans: (c)
By Chargaff's rules: A = T and C = G. Given A = 30% → T = 30%; given C = 20% → G = 20%.
2021
Q1: Complete the flow chart on central dogma (NEET 2021)
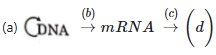
(a) (a) - Replication; (b) - Transcription;
(c) - Translation; (d) - Protein
(b) (a) - Transduction; (b) - Translation;
(c) - Replication; (d) - Protein
(c) (a) - Replication; (b) - Transcription;
(c) - Transduction; (d) - Protein
(d) (a) - Transcription; (b) - Replication;
(c) - Transcription; (d) - Transduction
Ans: (a)
Central dogma flow: DNA → (replication) DNA; DNA → (transcription) RNA; RNA → (translation) protein. Transduction refers to gene transfer by a virus and is unrelated to the central dogma steps listed.
Q2: What is the role of RNA polymerase III in the process of transcription in eukaryotes? (NEET 2021)
(a) Transcribes precursor of mRNA
(b) Transcribes only snRNAs
(c) Transcribes rRNAs (28S, 18S, and 5.8S)
(d) Transcribes tRNA, 5s rRNA and snRNA
Ans: (d)
- RNA pol III transcribes tRNA, 5S rRNA and some small RNAs (including some snRNAs). RNA pol I transcribes major rRNAs (28S, 18S, 5.8S). RNA pol II transcribes pre-mRNA (hnRNA).
Q3: DNA fingerprinting involves identifying differences in some specific regions in DNA sequence, called as (NEET 2021)
(a) Single nucleotides
(b) Polymorphic DNA
(c) Satellite DNA
(d) Repetitive DNA
Ans: (d)
- DNA fingerprinting relies on differences in repetitive DNA regions (like VNTRs, minisatellites and microsatellites). These repetitive sequences show high polymorphism among individuals and are used as markers.
Q4: Identify the correct statement. (NEET 2021)
(a) The coding strand in a transcription unit is copied to an mRNA.
(b) Split gene arrangement is characteristic of prokaryotes.
(c) In capping, methylguanosine triphosphate is added to the 3' end of hnRNA.
(d) RNA polymerase binds with the Rho factor to terminate the process of transcription in bacteria.
Ans: (d)
- Statement (d) is correct: Rho factor is involved in termination in many bacterial transcription units. (a) is incorrect because the template (non-coding) strand, not the coding strand, is copied. (b) is incorrect-split genes (introns/exons) are a eukaryotic feature. (c) is incorrect because the 5' cap (7-methylguanosine) is added at the 5' end; poly(A) tail is added at the 3' end.
Q5: If Adenine makes 30% of the DNA molecule, what will be the percentage of Thymine, Guanine and Cytosine in it? (NEET 2021)
(a) T:30 ; G:20 ; C:20
(b) T:20 ; G:25 ; C:25
(c) T:20 ; G:30 ; C:20
(d) T:20 ; G:20 ; C:30
Ans: (a)
Chargaff's rules: A = T, G = C. Given A = 30% → T = 30%. Remaining 40% is split equally between G and C → 20% each.
Q6: Which of the following RNAs is not required for the synthesis of protein? (NEET 2021)
(a) rRNA
(b) siRNA
(c) mRNA
(d) tRNA
Ans: (b)
- siRNA participates in gene silencing pathways (RNAi) and reduces specific mRNA levels; it is not a component required for normal translation. rRNA, mRNA and tRNA are essential components of the translation machinery.
Q7: Which is the "Only enzyme" that has "Capability to catalyze Initiation, Elongation, and Termination in the process of transcription in prokaryotes? (NEET 2021)
(a) DNA Ligase
(b) DNase
(c) DNA dependent DNA polymerase
(d) DNA dependent RNA polymerase
Ans: (d)
A single multi-subunit bacterial RNA polymerase carries out initiation (with sigma factor), elongation and termination (with factors such as rho) in prokaryotes.
Q8: Which one of the following statements about Histones is wrong? (NEET 2021)
(a) Histones are rich in amino acids - Lysine and Arginine.
(b) Histones carry a positive charge in the side chain.
(c) Histones are organized to form a unit of 8 molecules.
(d) The pH of histones is slightly acidic.
Ans: (d)
- Histones are basic proteins, rich in lysine and arginine, giving them a positive charge and basic pH. They assemble as an octamer (two each of H2A, H2B, H3 and H4).
Q9: Statement I: The codon 'AUG codes for methionine and phenylalanine.
Statement II: AAA' and 'AAG are both codons that code for the amino acid lysine.
In the light of the above statements, choose the correct answer from the options given below. (NEET 2021)
(a) Statement I is correct but Statement II is false
(b) Statement I is incorrect but Statement II is true
(c) Both Statement I and Statement II are true
(d) Both Statement I and Statement II are false
Ans: (b)
Statement I is false - AUG codes for methionine only (and serves as initiator codon). Statement II is true - both AAA and AAG code for lysine, illustrating degeneracy of the code.
2020
Q1: If the distance between two consecutive base pairs is 0.34 nm and the total number of base pairs of a DNA double helix in a typical mammalian cell is 6.6 × 109 dp, then the length of the DNA is approximately. (NEET 2020)
(a) 2.2 meters
(b) 2.7 meters
(c) 2.0 meters
(d) 2.5 meters
Ans: (a)
Length = number of bp × spacing = 6.6 × 109 × 0.34 × 10-9 m ≈ 2.244 m, so ~2.2 metres (option a).
Q2: The first phase of translation is: (NEET 2020)
(a) Aminoacylation of tRNA
(b) Recognition of an anti-codon
(c) Binding of mRNA to ribosome
(d) Recognition of DNA molecule
Ans: (a)
The first step is charging of tRNA with its cognate amino acid (aminoacylation) by aminoacyl-tRNA synthetases; this readies tRNAs for translation.
Q3: Name the enzyme that facilitates opening of DNA helix during transcription. (NEET 2020)
(a) DNA polymerase
(b) RNA polymerase
(c) DNA ligase
(d) DNA helicase
Ans: (b)
RNA polymerase binds to promoter and unwinds a short stretch of the DNA helix to expose the template strand for RNA synthesis during transcription initiation; hence RNA polymerase facilitates opening during transcription.
Q4: Which of the following statements is correct?
(a) Adenine does not pair with thymine
(b) Adenine pairs with thymine through one H-bond
(c) Adenine pairs with thymine through two H-bonds
(d) Adenine pairs with thymine through three H-bonds (NEET 2020)
Ans: (c)
In DNA, adenine pairs with thymine via two hydrogen bonds; guanine pairs with cytosine via three hydrogen bonds.
2019
Q1: Purines found both in DNA and RNA are (NEET 2019)
(a) Cytosine and thymine
(b) Adenine and thymine
(c) Adenine and guanine
(d) Guanine and cytosine
Ans: (c)
Purines are adenine and guanine, present in both DNA and RNA; pyrimidines include cytosine, thymine (DNA only) and uracil (RNA only).
Q2: Expressed Sequence Tags (ESTs) refers to (NEET 2019)
(a) Novel DNA sequences
(b) Genes expressed as RNA
(c) Polypeptide expression
(d) DNA polymorphism
Ans: (b)
ESTs are short cDNA sequences derived from mRNA and represent expressed genes (transcripts).
Q3: Match the following genes of the Lac operon with their respective products. (NEET 2019)
| (A) I gene | (i) β - galactosidase |
| (B) z gene | (ii) Permease |
| (C) a gene | (iii) Repressor |
| (D) y gene | (iv) Transacetylase |
Select the correct option.
| (A) | (B) | (C) | (D) | |
| (a) | (iii) | (iv) | (i) | (ii) |
| (b) | (i) | (iii) | (ii) | (iv) |
| (c) | (iii) | (i) | (ii) | (iv) |
| (d) | (iii) | (i) | (iv) | (ii) |
Ans: (d)
- I → repressor (iii); z → β-galactosidase (i); y → permease (ii); a → transacetylase (iv). Option (d) matches these roles.
Q4: Which of the following features of genetic code does allow bacteria to produce human insulin by recombinant DNA technology? (NEET 2019)
(a) Genetic code is specific.
(b) Genetic code is not ambiguous.
(c) Genetic code is redundant.
(d) Genetic code is nearly universal.
Ans: (d)
Because the genetic code is nearly universal, coding sequences from humans can be translated correctly in bacteria, enabling expression of human proteins (like insulin) in bacterial hosts.
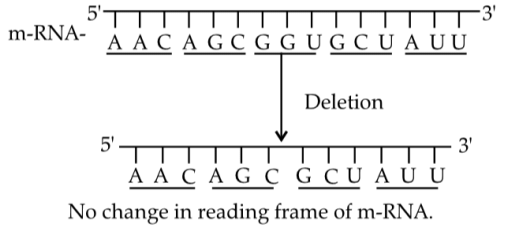
Q5: Under which of the following conditions there will be no change in the reading frame of following mRNA? (NEET 2019)
5' AACAGCGGUGCUAUU 3'
(a) Deletion of GGU from 7th, 8th and 9th positions
(b) Insertion of G at 5th position
(c) Deletion of G from 5th position
(d) Insertion of A and G at 4th and 5th position respectively
Ans: (a)
Deletion of three nucleotides (a codon) removes one amino acid but does not shift the reading frame. Any insertion/deletion of nucleotides not in multiples of three causes a frameshift.
2018
Q1: Select the correct match. (NEET 2018)
(a) Alec Jeffreys - Streptococcus pneumoniae
(b) Alfred Hershey and Martha Chase - TMV
(c) Matthew Meselson and F. Stahl - Pisum sativum
(d) Francois Jacob and Jacques Monod - Lac operon
Ans: (d)
Jacob & Monod proposed the lac operon model. Alec Jeffreys developed DNA fingerprinting; Meselson & Stahl demonstrated semi-conservative replication in E. coli; Hershey & Chase used bacteriophages to show DNA is genetic material.
Q2: The experimental proof for semi-conservative replication of DNA was first shown in a (NEET 2018)
(a) Fungus
(b) Bacterium
(c) Plant
(d) Virus
Ans: (b)
Meselson and Stahl demonstrated semi-conservative replication in the bacterium Escherichia coli.
Q3: Select the correct statement. (NEET 2018)
(a) Franklin Stahl coined the term '' linkage ''.
(b) Punnett square was developed by a British scientist.
(c) Spliceosomes take part in translation.
(d) Transduction was discovered by S. Altman.
Ans: (b)
- The Punnett square was developed by Reginald Punnett (British). Morgan coined the term 'linkage'. Spliceosomes are involved in RNA splicing (post-transcriptional), not translation. Transduction was discovered by Zinder & Lederberg.
Q4: Select the correct match. (NEET 2018)
(a) Ribozyme - Nucleic acid
(b) F2 x Recessive parent - Dihybrid cross
(c) T.H. Morgan - Transduction
(d) G. Mendel - Transformation
Ans: (a)
Ribozymes are catalytic RNAs (nucleic acids). Other matches are incorrect: Mendel - classical genetics, Morgan - linkage and fruit fly genetics, F2×recessive parent is test cross, not dihybrid cross.
Q5: Many ribosomes may associate with a single wiRNA to form multiple copies of a polypeptide simultaneously. Such strings of ribosomes are termed as (NEET 2018)
(a) Polysome
(b) Polyhedral bodies
(c) Plastidome
(d) Nucleosome
Ans: (a)
A polysome (polyribosome) is an mRNA molecule with multiple ribosomes attached, each synthesising a copy of the polypeptide simultaneously.
Q6: All of the following are part of an operon except (NEET 2018)
(a) An operator
(b) Structural genes
(c) An enhancer
(d) A promoter
Ans: (c)
Enhancers are eukaryotic regulatory elements and are not typical components of prokaryotic operons (which include promoter, operator and structural genes).
Q7: AGGTATCGCAT is a sequence from the coding strand of a gene. What will be the corresponding sequence of the transcribed mRNA? (NEET 2018)
(a) AGGUAUCGCAU
(b) UGGTUTCGCAT
(c) ACCUAUGCGAU
(d) UCCAUAGCGUA
Ans: (a)
The coding strand and mRNA are identical except that T in DNA is replaced by U in RNA. Thus AGGTATCGCAT (DNA coding, 5'→3') becomes AGGUAUCGCAU (mRNA).
2017
Q1: The final proof for DNA as the genetic material came from the experiments of (NEET 2017)
(a) Hershey and Chase
(b) Avery, MacLeod and McCarty
(c) Hargobind Khorana
(d) Griffith
Ans: (a)
Hershey and Chase (1952) provided final convincing evidence using radio-labelled bacteriophages showing DNA, not protein, carries genetic information.
Q2: If there are 999 bases in an RNA that code for a protein with 333 amino acids, and the base at position 901 is deleted such that the length of the RNA becomes 998 bases, how many codons will be altered? (NEET 2017)
(a) 11
(b) 33
(c) 333
(d) 1
Ans: (b)
Deletion at base position 901 affects that codon and all downstream codons. From 901 to 998 there are 98 bases → 98/3 ≈ 32.6 → 33 codons will be altered (including the one containing position 901).
Q3: During DNA replication, Okazaki fragments are used to elongate (NEET 2017)
(a) The lagging strand towards replication fork
(b) The leading strand away from replication fork
(c) The lagging strand away from the replication fork
(d) The leading strand towards replication fork
Ans: (c)
On the lagging strand synthesis is discontinuous: Okazaki fragments are synthesised in 5'→3' direction away from the replication fork, later joined to form a continuous strand.
Q4: Which of the following RNAs should be most abundant in animal cell? (NEET 2017)
(a) tRNA
(b) mRNA
(c) miRNA
(d) rRNA
Ans: (d)
rRNA makes up the bulk of cellular RNA (~80%) because ribosomes are numerous and rRNA is a major structural component.
Q5: Spliceosomes are not found in cells of (NEET 2017)
(a) Fungi
(b) Animals
(c) Bacteria
(d) Plants
Ans: (c)
Spliceosomes perform pre-mRNA splicing, a eukaryotic process. Prokaryotes (bacteria) generally lack introns and spliceosomal machinery.
Q6: The association of histone H1 with a nucleosome indicates that (NEET 2017)
(a) DNA replication is occurring
(b) The DNA is condensed into a chromatin fibre
(c) The DNA double helix is exposed
(d) Transcription is occurring
Ans: (b)
H1 binds to linker DNA and helps compact nucleosomes into higher-order chromatin fibres, indicating condensation rather than active transcription.
2016
Q1: Taylor conducted the experiments to prove semiconservative mode of chromosome replication on (NEET 2016 Phase 2)
(a) Vinca rosea
(b) Vicia faba
(c) Drosophila melanogaster
(d) E. coli
Ans: (b)
Taylor et al. (1957) used root-tip cells of Vicia faba labelled with tritiated thymidine to demonstrate semi-conservative replication at the chromosomal level.
Q2: The equivalent of a structural gene is (NEET 2016 Phase 2)
(a) Muton
(b) Cistron
(c) Operon
(d) Recon
Ans: (b)
The term cistron historically referred to a functional gene unit that codes for a polypeptide or functional RNA; it corresponds to a structural gene.
Q3: Which of the following rRNAs acts as structural RNA as well as ribozyme in bacteria? (NEET 2016 Phase 2)
(a) 5S rRNA
(b) 18S rRNA
(c) 23S rRNA
(d) 5.8S rRNA
Ans: (c)
In bacteria, the 23S rRNA (component of the large ribosomal subunit) has peptidyl transferase catalytic activity and acts as a ribozyme in peptide bond formation, besides being structural.
Q4: A molecule that can act as a genetic material must fulfill the traits given below, except (NEET 2016 Phase 2)
(a) It should be able to express itself in the form of 'Mendelian characters'
(b) It should be able to generate its replica
(c) It should be unstable structurally and chemically
(d) It should provide the scope for slow changes that are required for evolution.
Ans: (c)
Genetic material must be chemically stable to preserve information; it must be heritable, capable of replication, expressible and allow occasional changes for evolution. Structural instability would prevent reliable inheritance.
Q5: DNA-dependent RNA polymerase catalyses transcription on one strand of the DNA which is called the (NEET 2016 Phase 2)
(a) Template strand
(b) Coding strand
(c) Alpha strand
(d) Antistrand
Ans: (a)
The strand used by RNA polymerase as the template is the template (antisense) strand; it runs 3' → 5' and is complementary to the produced RNA.
Q6: Which one of the following is the starter codon? (NEET 2016 Phase 1)
(a) UAA
(b) UAG
(c) AUG
(d) UGA
Ans: (c)
AUG codes for methionine and functions as the initiator codon in most organisms (in bacteria it often codes for N-formylmethionine as the initiator).
Q7: Which of the following is required as inducer (s) for the expression of Lac operon? (NEET 2016 Phase 1)
(a) Lactose
(b) Lactose and Galactose
(c) Glucose
(d) Galactose
Ans: (a)
Lactose (or its isomer allolactose) acts as the physiological inducer that binds the lac repressor and allows the operon to be transcribed. Glucose represses lac operon via catabolite repression; galactose does not act as the inducer.
Q8: A complex of ribosomes attached to a single strand of RNA is known as (NEET 2016 Phase 1)
(a) Polypeptide
(b) Okazaki fragment
(c) Polysome
(d) Polymer
Ans: (c)
A polysome (polyribosome) is an mRNA associated with multiple ribosomes simultaneously translating it, producing many copies of the polypeptide.
FAQs on NEET Previous Year Questions(2016-25): Molecular Basis of Inheritance
| 1. What are the key differences between DNA and RNA that come up in NEET molecular basis of inheritance questions? |  |
| 2. How do Meselson and Stahl's experiment results prove semi-conservative DNA replication for NEET exams? |  |
| 3. What is the role of DNA polymerase and why does it matter for understanding molecular basis of inheritance? |  |
| 4. Why do students get confused between leading and lagging strand synthesis in NEET genetics problems? |  |
| 5. How do mutations during DNA replication affect inheritance traits tested in NEET exams? |  |







