Chapter Notes: Molecular Biology Techniques and Recent Advances in Molecular Biology - 2
DNA Sequencing Techniques
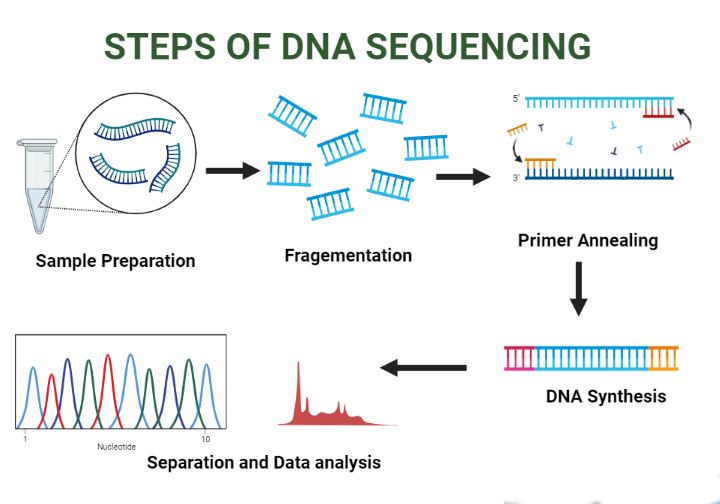
Sanger's Technique (Controlled Chain Termination Method)
- 2',3'-Dideoxynucleotides are used as chain terminators in this method.
- The DNA polymerase employed is Klenow polymerase.
- Sanger's method is capable of being automated, which enhances its efficiency.
- The incorporation of 2',3'-dideoxynucleotides halts DNA synthesis because these nucleotides lack a free 3' OH group necessary for forming the next phosphodiester bond.
Maxam and Gilbert's Chemical Cleavage Method
- This method, which predates contemporary sequencing techniques, involves...
Next Generation Sequencing (NGS)
- NGS encompasses a range of advanced DNA sequencing technologies that can generate vast amounts of sequence data simultaneously in a massively parallel fashion.
Comparison of Sanger's Sequencing and Next Generation Sequencing
- High Cost
- Low cost
- Single, simple homogeneous template DNA
- There are no specific requirements; a sample from almost any source can be used.
- It provides an "average" result for a DNA sample; however, results may be uninterpretable if the sample of DNA is heterogeneous.
- Well suited for heterogeneous DNA samples, as it can effectively analyse complex mixtures.
Three Basic Processes of NGS
Spatial separation
- At the beginning of the procedure, individual input DNA molecules are physically isolated from each other in space.
- The specifics of this process are platform-dependent.
Local Amplification
After the DNA molecules are separated, they are amplified in their original locations using a limited number of PCR cycles.
Parallel Sequencing
The amplified DNA molecules are sequenced simultaneously by adding polymerases and other necessary chemicals. Each distinct and amplified original molecule produces a "read" that corresponds to its specific sequence.
Some New Sequencing Techniques
- CAGE (Cap Analysis of Gene Expression). This method focuses on the selective capture, amplification, cloning, and sequencing of mRNAs through their 5' cap structure.
- RNA-Sequence. In this approach, mRNAs are converted into cDNAs using reverse transcription. These cDNAs are then amplified and sequenced directly.
- GRO-Sequence (Global Run-On Sequencing). This technique captures and sequences nascent transcripts specifically using NGS sequencing, helping to identify the locations of active transcription complexes.
- NET-Sequence (Native Elongating Transcript Sequencing). NET-Seq allows for the sequencing of RNA within elongating RNA polymerase-DNA-RNA complexes, facilitating genome-wide analysis of transcription in living cells.
- WES (Whole Exome Sequencing). Whole Exome Sequencing targets exons, which constitute about 1% of the human genome. This makes the exome a smaller and more manageable focus compared to the entire genome.
- Whole Exome Sequencing concentrates on exons, while whole genome sequencing encompasses the entire genome, including introns and regulatory regions. This distinction is crucial for diagnosing rare or hidden genetic diseases.
Genetically Modified Animals
Animals that have had foreign genes added to their fertilized eggs are called transgenic animals. These animals carry the introduced gene in every cell of their bodies and can pass it on to their offspring.
The gene of interest is typically a cloned piece of recombinant DNA that includes its own promoter, as well as an additional promoter that can be controlled.
Examples of Transgenic Animal Models
- Mus musculus (house mouse)
- Drosophila melanogaster (common fruit fly)
- Caenorhabditis elegans (nematode worm)
- Saccharomyces cerevisiae (baker's yeast)
- Escherichia coli (intestinal bacterium)
Applications of Transgenic Animals
- Study of DNA Regulatory Elements: Transgenic animals are used to study the regulatory elements of specific genes in various conditions. This helps researchers understand how genes are controlled in different situations.
- Examination of Gene Functions: These animals are useful for examining the functions of genes, particularly oncogenes, which are involved in the development of tumors. By studying transgenic animals, researchers can gain insights into how these genes contribute to cancer.
- Research on Gene Dosage Disorders: Transgenic animals are employed to investigate disorders that depend on gene dosage, such as overexpression or underexpression of certain genes. This research helps in understanding the impact of gene dosage on health and disease.
- Step Towards Gene Therapy: The creation of transgenic animals is a step towards gene therapy, which aims to treat genetic disorders by correcting or replacing faulty genes. Transgenic models provide a basis for developing and testing gene therapy approaches.
- Investigation of Physiological Effects: Transgenic animals allow scientists to investigate the physiological effects of adding or removing a specific gene from the genome. This research helps in understanding the functions and roles of individual genes in health and disease.
- Unique Genetic Models: Transgenic animals provide unique genetic models for studying pathology and pharmacology. These models help researchers understand disease mechanisms and test potential treatments in a controlled genetic background.
Various Approaches to Genetic Modification
- Injection: Involves injecting the transgene into the male pronucleus of a fertilized egg.
- Homologous Recombination: This process occurs in embryonic stem cells and is known as targeted mutagenesis.
- Forward Genetics:. strategy used in genetic research.
- Animal Cloning:. method used to create genetically identical animals.
Fundamental Concept of Targeted Mutagenesis
- Targeted mutagenesis involves replacing a normal gene with a neo-disrupted recombinant gene in a small percentage of embryonic stem cells through homologous recombination.
- There are different forms of targeted mutagenesis:
- Gene Knockout: Replaces an endogenous gene with a mutated transgene in embryonic stem cells through homologous recombination.
- Gene Knock-in: Swaps a mutated endogenous gene with a normal transgene in embryonic stem cells via homologous recombination.
- Gene Knockdown: Involves gene silencing induced by siRNA or miRNA, known as RNA interference (RNAi).
- Gene Knockup: Increases gene transcription using transcription factors.
Procedure for Creating a Knockout Mouse
- The first step is to inactivate the specific gene that we want to study.
- Next, embryonic stem (ES) cells are cultured from mice.
- These ES cells are then transfected with a cloned mutated non-functional gene.
- Only a small number of these cultured embryonic cells will incorporate the non-functional gene through a process called homologous recombination.
- The ES cells with the altered gene are then isolated.
- These altered ES cells are microinjected into a mouse blastocyst.
- The blastocyst is then implanted into a foster mother.
- Finally, the mice are bred for several generations until offspring are produced that have the altered gene in their germ cells.
Characteristic Features of Different Strategies of Transgenic Models
1. Transgenic: Pronuclear Injection of Transgene
- This method is widely used for creating transgenic models.
- It involves the random integration of the transgene into the host genome.
- Genomic DNA or cDNA constructs can be injected into the pronucleus.
- There is variable expression of the transgene in each individual founder animal.
- Gain-of-function models are created by overexpressing the transgene using tissue-specific promoters.
- Loss-of-function models are generated using antisense or dominant negative transgenes.
- Inducible expression of the transgene is also possible with this method.
- This technique is applicable to various species, including mice, rats, and zebrafish.
Targeted Mutagenesis
- Site-specific integration of transgene
- Primarily used in mice
- Characteristics of gene knockout:
- Tissue-specific knockout is possible
- Absence of a phenotype may occur due to redundancy, not solely because of the knockout
- Characteristics of gene knock-in:
- Can accurately model human disease
- Latest Techniques:
- miRNA/siRNA-mediated gene silencing: Also known as gene knockdown, this technique involves using microRNA or small interfering RNA to silence specific genes, reducing their expression.
- CRISPR-Cas9-mediated gene editing: This powerful tool can create both loss-of-function and gain-of-function mutations in genes, allowing precise modifications to the genome.
- Random mutations with ENU: N-ethyl-N-nitrosourea (ENU) induces random mutations in the genome. A specific phenotype is selected, and then genetic characterization follows to identify novel genes associated with that phenotype.
- Nuclear Transfer:
- Enucleation: The nucleus of an oocyte (egg cell) is removed, creating an enucleated oocyte.
- Fusion: The enucleated oocyte is fused with the nucleus of a somatic cell (a non-reproductive cell), transferring the genetic material into the oocyte.
- Implantation: The fused cell, now containing the somatic nucleus, is implanted into the uterus of a surrogate mother, where it can develop into an embryo.
Characteristics of Animal Cloning
- Successful in various mammalian species including sheep, mice, cows, and monkeys
- Cloning of genetically identical individuals is feasible
- May impact lifespan
- Numerous ethical concerns exist
Successful Stories of Animal Cloning
- In 1996, Dolly the sheep was cloned by Ian Wilmut and Keith Campbell, making history as the first cloned mammal from an adult somatic cell.
- Bonnie was the first lamb born to Dolly, marking a significant milestone in the field of animal cloning.
Antisense Nucleic Acid in Research and Therapy
- Antisense nucleic acid can be either RNA or DNA.
- It can be natural or synthetic.
- This nucleic acid is complementary to mRNA.
- siRNA can be utilized.
- Antisense nucleic acid bonds with target mRNA, leading to its selective dissolution or inhibition.
- Gene knockdown techniques are employed to examine the role of specific genes.
- These nucleic acids can be applied in therapeutic strategies for viral infections like HIV.
Hybridoma
- The hybridoma technique is used to produce monoclonal antibodies in a clinical laboratory setting.
- This method was first introduced by Georges Köhler and César Milstein in 1975.
- Monoclonal antibodies are designed to target a specific epitope of an antigen.
- In this process, spleen cells from immunised animals are fused with mouse myeloma cells to create hybrid cells.
- Polyethylene Glycol (PEG - 1500) is used as the fusing agent in this process.
- The hybrid cells are then grown in HAT medium, which contains Hypoxanthine, Aminopterine, and Thymidine.
- Unfused normal cells are unable to survive in this environment because they lack the ability to multiply.
Death of Unfused Myeloma Cells
- Aminopterine, a folate antagonist, inhibits the synthesis of new purines and DNA in unfused myeloma cells because these cells lack the enzyme HGPRTase.
- Normal cells produce the HGPRTase enzyme, allowing them to survive alongside myeloma cells.
- Myeloma cells possess the ability to replicate and multiply.
- Positive clones are selected and their numbers are increased.
- A large quantity of antibodies can be harvested, which is crucial for the production of monoclonal antibodies.
- Counting lymphocyte subpopulations is essential for understanding the immune system's response.
- Specific cells are prepared in a quantitative manner.
- A nephelometric assay is used to analyze blood components.
- This method aids in the preparation of monoclonal antibodies for ELISA tests.
- Quantitative preparation of pure antigens is also carried out.
Restriction Fragment Length Polymorphism (RFLP)
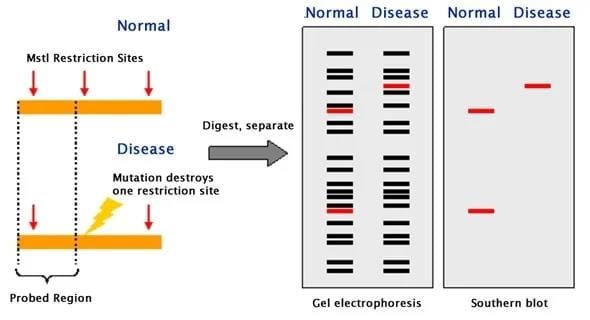
RFLP is a term used to describe genetic differences based on how restriction endonucleases cut DNA. These differences occur when there is a change in the DNA that either creates a new cutting site or removes an existing one, leading to variations in the patterns produced by these enzymes.
There are two main types of DNA variations that can result in RFLP:
- Variable Number Tandem Repeat (VNTR)
RFLP due to Mutations Altering Restriction Sites
RFLP (Restriction Fragment Length Polymorphism) can occur when certain mutations change the restriction site in the DNA. A well-known example is sickle cell anaemia, which is caused by a point mutation. This mutation alters the DNA sequence in such a way that it removes the recognition site for the restriction endonuclease MstII.
Procedure of RFLP
- DNA is extracted from the cell.
- The DNA is cut by a specific restriction endonuclease.
- The resulting DNA fragments are separated using agarose gel electrophoresis.
- DNA fragments are denatured.
- The DNA is transferred to a nitrocellulose membrane using the Southern blot technique.
- It is treated with a radiolabelled probe.
- DNA variation is detected by checking for hybridisation through autoradiography.
Applications of RFLP
- Tracing chromosomes from parents to offspring and conducting prenatal diagnosis for diseases.
- Direct diagnosis of sickle cell disease using RFLP.
- Indirect, prenatal diagnosis of phenylketonuria.
Medicolegal Applications
- Detecting mutations, including point mutations, insertions, and deletions.
DNA Fingerprinting
- DNA fingerprinting involves using normal genetic variations such as SNPs, VNTRs, or RFLPs to generate a unique pattern of DNA fragments for each individual.
- This technique is also referred to as DNA profiling.
- VNTR and RFLP are commonly used alongside SNP in DNA profiling.
- The concept of DNA fingerprinting was pioneered by Alec Jeffreys in 1985.
DNA Footprinting
- DNA that is bound to a protein remains intact and is not degraded by DNase enzymes.
- During sequencing of this type of DNA, a protected region is observed, indicating the 'footprint' of the bound protein.
- This phenomenon occurs because nucleases are unable to cleave the DNA that is directly associated with the protein.
Chromosome Walking
- Chromosome walking is a technique used to isolate and clone target DNA from a large DNA segment.
- In this method, a fragment from one end of a long DNA piece is used to identify another overlapping fragment that extends beyond the first.
- This process is repeated iteratively until the target DNA is successfully retrieved.
- One of the early successes of chromosome walking was the isolation of the cystic fibrosis (CF) gene, highlighting the effectiveness of this technique.
Chromosome Walking
- To isolate Gene X from a segment of DNA, we start with a probe that targets a section of DNA at the 5' end.
- The exact position of Gene X is unknown at this stage.
- The probe attaches to the first fragment of DNA, which is a step towards locating Gene X.
- By using the same probe, we continue to search for subsequent fragments, walking along the chromosome until we find the fragment containing Gene X.
Chromosome Jumping
- Chromosome jumping is a technique used to bypass regions of DNA that are challenging to clone.
- These challenging areas often include regions with repetitive DNA, which complicate the mapping process using chromosome walking.
- Repetitive DNA consists of sequences that are present multiple times in the genome, making cloning efforts more difficult.
Understanding Stem Cell Biology
What are Stem Cells?
- Stem cells are defined by their unique abilities: self-renewal, which is the capacity to produce identical daughter cells, and potency, the potential to differentiate into various specialized cell types.
Types of Stem Cells
1. Totipotent Cells
- Totipotent cells have the remarkable ability to develop into an entire organism on their own.
- This capability is typically seen in a fertilized egg, known as a zygote.
2. Pluripotent Cells
- Pluripotent cells can differentiate into nearly all cell types in the body, including those from the endoderm, mesoderm, and ectoderm germ layers.
- Examples include embryonic stem (ES) cells, which are a specific type of pluripotent stem cell.
Multipotent cells
- These cells have the ability to develop into multiple types of cells, but they cannot create all the cell types found in the body.
- An example of multipotent cells is Hematopoietic Stem (HS) cells, which can give rise to various blood cell types.
Oligopotent cells
- Oligopotent cells are also referred to as progenitor or precursor cells.
- These cells have the capacity to form more than one type of cell lineage.
- An example of oligopotent cells is Neural Stem cells, which can differentiate into various types of neural cells.
Unipotent cells
- Unipotent cells have the ability to develop into a single type of cell lineage.
- An example of unipotent cells is spermatogonial stem (SS) cells, which give rise to sperm cells.
Nuclear Reprogramming
This is the process of reverting specialized cells back to a totipotent or pluripotent state, where they can develop into any cell type.
Transdifferentiation
- Transdifferentiation involves changing multipotent cells that are committed to a specific lineage into different cell types beyond their usual specialization.
Stem cells are unique cells in the body that have the ability to develop into different types of cells. They are like blank slates that can take on various roles in the body. There are different types of stem cells, each with its own characteristics and potential.
Types of Stem Cells
1. Embryonic Stem Cells (ES, ESC)
- Source: Inner cell mass from blastocysts.
- Properties: Maintain a stable euploid karyotype after extensive culture and can become various cell types in vitro.
2. Embryonic Germ Cells (EG, EGC)
- Source: Primordial germ cells from embryos.
- Properties: Similar pluripotency to ES cells.
3. Trophoblast Stem Cells (TS, TSC)
- Source: Trophoblast layer.
- Properties: Can form all trophoblast subtypes when injected into blastocysts.
4. Extra Embryonic Endoderm Cells (XEN)
- Source: Inner cell mass from blastocysts.
- Properties: Contribute only to the parietal endoderm lineage when injected into a blastocyst.
5. Embryonal Carcinoma Cells (EC)
- Source: Teratocarcinoma.
- Properties: Can develop into nearly all cell types when injected into blastocysts.
6. Mesenchymal Stem Cells (MS, MSC)
- Source: Bone marrow, muscle, adipose tissue, peripheral blood, umbilical cord blood.
- Properties: Differentiate into mesenchymal cell types, such as adipocytes, osteocytes, chondrocytes, and myocytes.
7. Multipotent Adult Stem Cells (MAPC)
- Source: Mononuclear cells from bone marrow.
- Properties: Can differentiate into all tissues in vivo when injected into a mouse blastocyst.
8. Spermatogonial Stem Cells (SS, SSC)
- Source: Newborn testis.
- Properties: Reconstitute long-term spermatogenesis after transplantation and restore fertility.
9. Germline Stem Cells (GS, GSC)
- Source: Neonatal testis.
- Properties: Contribute to the formation of germ layers and various tissues when injected into blastocysts.
10. Multipotent Adult Germline Stem Cells (maGSC)
- Source: Adult testis.
- Properties: Differentiate into three germ layers in vitro and contribute to various tissues when injected into blastocysts.
11. Neural Stem Cells (NS, NSC)
- Source: Fetal and adult brain (subventricular zone, ventricular zone, hippocampus).
- Properties: Can differentiate into neurons and glial cells both in vivo and in vitro.
12. Unrestricted Somatic Stem Cells (USSC)
- Source: Mononuclear fraction of cord blood.
- Properties: Differentiate into various cell types in vitro and contribute to various cell types in transplantation.
13. Epiblast Cells (EpiSC)
- Source: Early postimplantation epiblast.
- Properties: Differentiate into three germ layers in vitro but do not contribute to normal tissues when injected into blastocysts.
14. Induced Pluripotent Stem Cells (iPS, iPSC)
- Source: Various terminally differentiated cells and tissue stem cells.
- Properties: Can be created from different somatic cell types using various transcription factors and small molecules.
Gene Therapy
Gene therapy is an innovative approach in the field of therapeutics that involves the intracellular delivery of genes to address and rectify existing health issues at the genetic level. The aim is to create a therapeutic effect by fixing problems within the cells.
Gene therapy can be categorized into two main types:
- Somatic cell gene therapy. This involves introducing a therapeutic gene into somatic cells, which are the non-reproductive cells in the body. The goal is to correct genetic defects or introduce beneficial genes to treat diseases.
- Germ cell gene therapy. In this type, the gene is altered in germ cells, which are the reproductive cells. This can lead to heritable changes, meaning the genetic modifications can be passed on to future generations. This is different from transgenic animals, which are created through such therapies.
Methods of Gene Delivery
Chemical Methods of Gene Delivery: DEAE-Dextran
- DEAE-Dextran, short for Diethylaminoethyl-dextran, is a carbohydrate polymer known as dextran.
- This polymer carries a positive charge, enabling it to bind to the negatively charged backbone of DNA.
- The complex formed between DEAE-Dextran and DNA retains a positive charge, allowing it to interact with the negatively charged surfaces of cell membranes.
- The cellular uptake of this complex occurs likely through a process called endocytosis, where the cell membrane engulfs the complex and brings it into the cell.
Calcium Phosphate
- Mix DNA with calcium chloride.
- Add this mixture to a phosphate buffered saline solution.
- Incubate at room temperature.
- A DNA-containing precipitate forms.
- This precipitate is applied to cultured cells.
- Cells take in the precipitate through endocytosis or phagocytosis.
- Simplicity. The method is straightforward and easy to perform.
- Applicability. It can be used on a wide range of cell types.
- Stability. It is effective in generating stably transfected cell lines, which are important for long-term studies of gene expression.
Cationic Lipids (Lipofection)
- Liposomes and DNA: When lipids are combined with DNA in water, they form hollow spheres called liposomes, with DNA trapped inside.
- Cell Entry: When these liposomes are introduced to cells in a lab setting (in vitro), some of them merge with the cell membranes and enter the cells through a process known as endocytosis.
Polymers
- Transfection with Polymers: Recently, various organic polymers have been utilized for transfection purposes.
- Polyethylenimine (PEI): One of the most common polymers used is a polycation called polyethylenimine (PEI). PEI is a large organic molecule with a high positive charge density, often referred to as a "proton sponge."
- Mechanism of Action: PEI compacts DNA into positively charged particles that can interact with negatively charged cell surfaces. This interaction facilitates the entry of DNA into cells via endocytosis.
Microinjection
- Microinjection is a technique where DNA is directly injected into the nuclei of target cells using fine glass needles while observing under a microscope.
- Although the concept is simple, microinjection is one of the most difficult methods to execute.
- The process is tedious and time-consuming, typically allowing only a few hundred cells to be transfected in each experiment.
Electroporation
- Electroporation is a method used to introduce nucleic acids into cells by subjecting them to a rapid pulse of high-voltage current.
- This high-voltage pulse temporarily creates pores in the cell membrane, facilitating the entry of external DNA into the cytoplasm.
- However, the efficiency of gene transfer through electroporation is generally low, and the method often results in a high rate of cell death.
Gene Gun
- In the gene gun method, plasmid DNA is coated onto metal microparticles and then shot into cells using either electrostatic force or gas pressure.
- Some DNA becomes trapped in cells and may be expressed at sufficient levels.
- This technique is fast, simple, and safe, and it can transfer genes to a wide variety of tissues.
- However, there are known limitations related to the size of the plasmid DNA and the capacity of vectors used for gene delivery.
Viral Vectors in Gene Delivery
- Viral vectors have the main advantages of persistent gene transfer in dividing cells.
- The main disadvantages include the theoretical risk of insertional mutagenesis.
- Retroviral:
- Persistent gene transfer in dividing cells.
- Theoretical risk of insertional mutagenesis.
- Lentiviral:
- Persistent gene transfer in transduced tissues.
- Might induce oncogenesis in some cases.
- Adenoviral:
- Highly effective in transducing various tissues.
- Viral capsid elicits strong immune responses.
- Adeno-Associated viral:
- Elicits few inflammatory responses, nonpathogenic.
- Limited packaging capacity.
Human foamy virus:
- Persistent gene expression in both dividing and non-dividing cells.
- Needs a stable packaging system.
HSV-1:
- Large packaging capacity with persistent gene transfer.
- Residual cytotoxicity with neuron specificity.
Simian virus-40:
- Wide host cell range; lack of immunogenicity.
Alpha viruses:
- Limited immune responses against the vector.
- Transduced gene expression is transient.
Nonviral Vectors
Transposon/Transposase system:
- Undetermined, probably large packaging capacity.
- Transfects many cell types with long-term gene expression.
- Early stage in development.
Liposome:
- Transfects many cell types.
- Large holding capacity to enable a high number of base pairs.
- Expensive to produce.
Naked DNA:
- Efficient in gene transfer.
- Limited immunogenicity.
- Transient and low-level expression.
Site Specific Integrase:
- Specific integration site.
Strategies of Gene Transfer
- In vivo gene therapy involves directly transferring genes into the patient.
- Ex vivo refers to the process where cells are removed from the patient for gene transfer.
- Gene transfer occurs into target cells.
- Target cells are returned to the patient after gene transfer.
Application of Gene Therapy in Medicine
- The following are the primary areas where gene therapy clinical trials have focused:
- Cancer (67%)-Most common area of focus.
- Vascular Disease (8.9%)
- Monogenic Disorders (8.6%)
Gene Therapy in Cancers
Local/Regional Approach
Suicide Gene/Prodrug:
- Intratumoral injection of an adenoviral vector expressing the thymidine kinase (TK) gene.
- Cells that take up and express the TK gene are sensitive to Ganciclovir.
When is this Treatment Recommended?
- This treatment works well for tumours that come backin the same place, specifically in the:
- Prostate
- Breas
- Colon
- Also used for Glioblastoma Multiforme (a type of brain cancer)
Tumor Suppressor Gene Therapy
- Involves using adenoviruses to deliver the p53 tumor suppressor gene to cancer cells.
- This approach is being explored for treating:
- Squamous cell carcinoma of the head and neck
- Esophageal cancer
- Non-small cell lung cancer
Oncolytic Virus Therapy
- This therapy uses viruses that are engineered to replicate only in tumour cells and not in normal cells.
- It is being investigated for treating squamous cell carcinoma of the head and neck.
Systemic Gene Therapy
- Further details on systemic gene therapy will be provided in the future.
Immunomodulation
- Genes that enhance the immune system include those responsible for producing cytokines, chemokines, or co-stimulatory molecules.
Anti-Angiogenesis
- Continuous production of angiogenesis inhibitors such as angiostatin and endostatin.
- Using siRNA to reduce levels of vascular endothelial growth factor (VEGF) or its receptor.
Gene Therapy for Vascular Disease
- Transgenes for vascular endothelial growth factor (VEGF), fibroblast growth factor (FGF), and hypoxia-inducible factor that improve blood flow.
- Applications in skeletal muscle for critical limb ischemia.
- Applications in cardiac muscle for angina or myocardial ischemia.
Gene Therapy for Genetic Disorders
- Gene transfer approaches for treating genetic disorders primarily focus on gene addition therapy.
- This technique involves inserting a missing gene into a target cell that plays a crucial role in the body.
- The first genetic disorder successfully treated with gene therapy was Severe Combined Immunodeficiency (SCID).
- Dr. W. French Anderson is renowned as the "Father of Gene Therapy."
Gene Therapy for Other Diseases
- In the case of Parkinson's disease, adeno-associated virus (AAV) vectors are employed to deliver genes that produce enzymes essential for increasing levels of dopamine or the inhibitory neurotransmitter Gamma-aminobutyric acid (GABA).
- For Alzheimer's disease, fibroblasts are modified using a retroviral vector to express nerve growth factor (NGF), which supports the survival and growth of neurons.
Introduction to the Human Genome Project
The Human Genome Project, launched in 1990 by the United States, was a groundbreaking endeavor aimed at mapping the entire DNA sequence of the human genome. This ambitious project involved the development of automated techniques, advanced instruments, and specialized software for analyzing the complex genetic code that makes up human DNA.
Bioinformatics
Bioinformatics is a crucial field that deals with the collection, storage, and analysis of biological data, particularly focusing on DNA and protein sequences. It plays a vital role in understanding the genetic and proteomic information of organisms.
Key Concepts in Bioinformatics
- Genome: The complete set of genes present in an organism, which carries the genetic information necessary for its growth, development, and functioning.
- Genomics: The comprehensive study of the structure and function of genomes, including the analysis of gene sequences, their roles, and interactions within the genome.
- Transcriptome: The entire collection of RNA transcripts produced by the genome within a specific timeframe, reflecting the genes that are actively expressed at that time.
- Transcriptomics: The in-depth study of the transcriptome, focusing on the analysis of RNA transcripts to understand gene expression patterns and regulatory mechanisms.
- Proteome: The complete set of proteins present in an organism, which are essential for various biological functions and processes.
- Proteomics: The systematic investigation of the structures, functions, and variations of proteomes, particularly in the context of health and disease.
- Glycome: The entire range of simple and complex carbohydrates, known as glycans, present in an organism.
- Glycomics: The comprehensive study of glycomes, such as the human glycome, to understand the roles and functions of glycans in biological systems.
- Lipidome: The complete set of lipids, including fats and oils, found in an organism.
- Lipidomics: The detailed study of lipid structures, functions, and interactions, particularly in relation to health and disease conditions.
- Metabolome: The entire collection of metabolites, which are small molecules involved in metabolic processes, present in an organism.
- Metabolomics: The thorough investigation of metabolite structures, functions, and variations across different metabolic states and conditions.
- Nutrigenomics: The systematic study of the impact of nutrients on gene expression and how genetic variations influence nutrient metabolism and utilization.
- Pharmacogenomics: The application of genomic information and technologies to improve drug discovery, development, and personalized medicine approaches.
- Exome: The sequence of all mRNA exons, which are the coding regions of genes, expressed in a specific cell, tissue, organ, or organism, providing insights into the protein-coding potential of the genome.
First Generation Proteomics
- SDS PAGE or 2D Electrophoresis are used to purify protein samples.
- To determine the amino acid sequence, end-group analysis methods like Sanger's reagent and Edman's reagent are employed.
Second Generation Proteomics
- Nanoscale chromatographic techniques are utilized to purify proteins in the sample.
- To determine the amino acid sequence, methods such as Mass Spectrometry and MudPIT (Multidimensional Protein Identification Technology) are used.
- Successive rounds of chromatography separate peptides from complex biological samples into simpler fractions for individual analysis by Mass Spectrometry.
Bioinformatic and Genomic Resources
These databases are crucial for supporting various types of research, including molecular, biochemical, epidemiological, and clinical studies.
UniProt Knowledgebase (UniProtKB)
- The UniProt Knowledgebase (UniProtKB) is a joint initiative by the Swiss Institute of Bioinformatics and the European Bioinformatics Institute.
- Its goal is to provide the scientific community with a comprehensive, high-quality, and freely accessible resource for protein sequence and structural information.
UniProtKB is divided into two main sections:
- Swiss-Prot. This section contains entries where the functions, domain structures, and post-translational modifications of proteins have been confirmed through manual curation.
- TrEMBL. This section includes protein sequences derived from genomes, where potential functions are assigned automatically using computer algorithms.
In summary, UniProtKB is a vital resource for researchers studying protein sequences and structures, with Swiss-Prot offering manually curated data and TrEMBL providing computationally annotated sequences.
Gen Bank
GenBank is a database that compiles and stores all known biological nucleotide sequences along with their corresponding translations, making this information easily searchable.
PDB
- The Protein Data Bank (PDB) is a repository of three-dimensional structures of proteins, polynucleotides, and other biological macromolecules.
- As of now, the PDB contains over 95,000 three-dimensional structures of proteins.
Tagged SNPs
- When groups of Single Nucleotide Polymorphisms (SNPs) on the same chromosome are inherited together, they form blocks known as haplotypes.
- Tag SNPs are specific SNPs within these haplotype blocks that act as unique markers for the entire block.
- These tag SNPs help identify and study specific genetic variations associated with particular diseases or physiological responses.
HapMap
The HapMap Project, short for the Haplotype Map Project, is a significant research effort aimed at identifying Single Nucleotide Polymorphisms (SNPs) associated with common human diseases and varying responses to medications.
The primary goal of this project is to facilitate earlier and more accurate diagnoses of potential genetic risks, which can ultimately lead to improved prevention strategies and more effective patient care.
Encode
The ENCODE (Encyclopedia of DNA Elements) Project was initiated by the National Human Genome Research Institute (NHGRI). This project is a collaborative effort that combines laboratory and computational approaches to identify every functional element within the human genome.
ENCODE focuses on various aspects of the genome, including:
- Mapping sites of DNA methylation, which involves studying chemical modifications to DNA that can affect gene expression.
- Assessing local histone methylation, a process that plays a crucial role in the regulation of gene activity by modifying histone proteins around which DNA is wrapped.
- Investigating other related biological processes that contribute to our understanding of genome function.
Entrez Gene
- Entrez Gene is a database operated by the National Center for Biotechnology Information (NCBI) that offers a range of information about individual human genes.
- The database includes details such as:
- Genomic sequences
- Exon-intron boundaries
- mRNA sequences generated from the genes
- Known phenotypes associated with mutations in the genes
dbGAP
- dbGAP, which stands for the Database of Genotype and Phenotype, is another resource provided by NCBI that complements Entrez Gene.
- This database focuses on collecting research findings related to the relationships between specific genotypes (genetic makeup) and phenotypes (observable traits).
Linkage And Association Studies
- Genes that are close together on a chromosome are usually inherited together, unless they are separated by a process called recombination.
- When two genes are nearby each other on the chromosome, the likelihood of them being separated during recombination is low.
Linkage and Association Studies
- There are two primary approaches for mapping genes associated with human diseases:
- Classic linkage analysis can be performed using a known genetic model.
- If the genetic model is unknown, researchers can study pairs of affected relatives to identify potential links.
- Disease genes can also be mapped through allelic association studies, which encompass Genome Wide Association Studies (GWAS).
Linkage Analysis
- Linkage disequilibrium refers to the phenomenon where genetic marker loci that are in close proximity to a disease allele are inherited together within families.
- This occurs because these markers are genetically linked to the disease allele, making them useful for tracking the inheritance of the allele.
- In linkage analysis, researchers evaluate genetic marker loci in family members who exhibit the disease or trait of interest.
- Over time, a set of marker loci that co-segregate with the suspected disease can be identified.
- This process helps in locating and cloning the disease allele.
- The genetic markers commonly used in linkage analysis include single nucleotide polymorphisms (SNPs) and repeat-length polymorphisms, such as minisatellite and microsatellite repeats.
- In a Genome-Wide Association Study (GWAS), researchers analyze large groups of patients with and without a disease across the entire genome to identify genetic variants or polymorphisms that are more prevalent in patients with the disease.
- This approach helps pinpoint regions of the genome that contain a variant gene or genes associated with increased disease susceptibility.
- Within these identified areas, potential causal variants are tentatively identified using a "candidate gene" approach.
- This involves selecting genes based on their strong association with the disease and their likely biological function related to the disease.
- By doing so, researchers aim to locate the causal gene or clarify the functional polymorphism linked to the disease.
Comparison of Linkage and Association Studies
- Linkage Study
- Association Study
- Conducted within the same family or among sibling groups
- Compares a group of affected individuals with a control group
- Linkage studies are generally more effective for analysing monogenic traits
- Suitable for identifying susceptibility genes related to polygenic and multifactorial disorders
- Obtaining sufficient statistical power for studying complex traits can be challenging
- Linkage studies offer greater statistical power for detecting complex multigenic disorders
FAQs on Chapter Notes: Molecular Biology Techniques and Recent Advances in Molecular Biology - 2
| 1. What are the main DNA sequencing techniques used in molecular biology? |  |
| 2. How are genetically modified animals created and what are their applications? |  |
| 3. What is a hybridoma and how is it used in research? |  |
| 4. What is Restriction Fragment Length Polymorphism (RFLP) and how is it applied in genetics? |  |
| 5. What is gene therapy and what are its potential benefits? |  |






