Practice Questions :Gene Expression
SECTION I: MULTIPLE CHOICE
Directions
Answer the following 20 questions. Each question has four answer choices (A, B, C, D). Select the one best answer for each question unless otherwise directed. Some questions may instruct you to select TWO answers; these will be clearly marked.
Calculators are not permitted on this section.
Questions 1-3 refer to the following experimental data.
Figure 1: Effect of Lactose on lacZ Gene Expression
Researchers measured β-galactosidase enzyme activity (the product of the lacZ gene) in E. coli cultures grown under different conditions. Results are shown below:
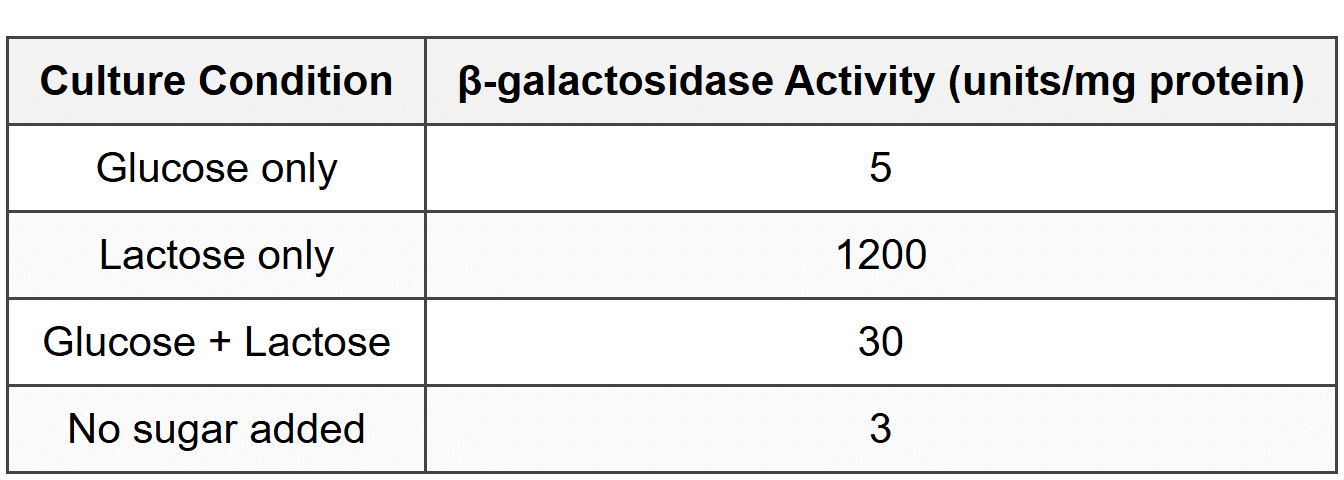

1. Which of the following best explains why β-galactosidase activity is low when both glucose and lactose are present?
- Glucose competitively inhibits the active site of β-galactosidase enzyme.
- The presence of glucose prevents CAP-cAMP complex formation, reducing transcription of the lac operon even when lactose is present.
- Lactose cannot enter the cell when glucose is present due to transport protein competition.
- Glucose causes the lac repressor to bind more tightly to the operator sequence.
2. In the "Lactose only" condition, which of the following molecular events occurs at the lac operon?
- Allolactose binds to the repressor protein, causing it to release from the operator and allowing transcription to proceed.
- Lactose binds directly to RNA polymerase, increasing its affinity for the promoter region.
- The repressor protein is degraded by proteases activated by lactose.
- Lactose methylates the promoter region, preventing repressor binding.
3. A mutation in the lacI gene results in a repressor protein that cannot bind to the operator. Based on the data in Figure 1, predict the β-galactosidase activity in a mutant strain grown in "Glucose only" conditions.
- Activity would remain at approximately 5 units/mg protein because glucose still prevents CAP-cAMP activation.
- Activity would increase to approximately 1200 units/mg protein because the operon cannot be repressed.
- Activity would increase to approximately 30 units/mg protein, matching the Glucose + Lactose condition.
- Activity would decrease to 0 units/mg protein because the mutation prevents all transcription.
Questions 4-5 refer to the following diagram.
Figure 2: Eukaryotic Gene Structure and Processing
The diagram below represents a eukaryotic gene and its processing pathway:
DNA: [Promoter]--[Exon 1]--[Intron 1]--[Exon 2]--[Intron 2]--[Exon 3] Pre-mRNA: [Exon 1]--[Intron 1]--[Exon 2]--[Intron 2]--[Exon 3]--[polyA tail] ↓ Mature mRNA: [Exon 1]--[Exon 2]--[Exon 3]--[polyA tail]
4. Which statement accurately describes a key difference between prokaryotic and eukaryotic gene expression illustrated by this diagram?
- Prokaryotic genes contain introns that are removed by spliceosomes during transcription.
- Eukaryotic genes undergo RNA processing including splicing, 5' capping, and polyadenylation before translation.
- Prokaryotic mRNA molecules receive a poly-A tail but lack a 5' cap structure.
- Eukaryotic transcription and translation occur simultaneously in the cytoplasm.
5. Select TWO answers. Alternative splicing of the pre-mRNA shown in Figure 2 could produce different mature mRNA molecules. Which TWO of the following represent accurate consequences of alternative splicing?
- Different combinations of exons in the mature mRNA can produce proteins with different amino acid sequences.
- Alternative splicing increases the coding capacity of the genome by allowing one gene to produce multiple protein variants.
- Alternative splicing always results in nonfunctional proteins due to frameshift mutations.
- The process of alternative splicing occurs in the nucleus and requires ribosomal subunits.
6. A researcher adds actinomycin D, a drug that blocks RNA polymerase, to a culture of actively dividing eukaryotic cells. Three hours later, the researcher observes that protein synthesis continues in these cells, though at declining levels. Which of the following best explains this observation?
- Cells can synthesize proteins without mRNA by using DNA as a direct template for translation.
- Existing mRNA molecules in the cytoplasm continue to be translated until they are degraded.
- Ribosomes can continue to function independently of mRNA for several hours after transcription stops.
- The drug does not affect transcription in eukaryotes because RNA polymerase is located in the nucleus.
Questions 7-8 refer to the following information.
Experimental Scenario: Studying Transcription Factors
Scientists are studying a transcription factor protein (TF-X) that regulates the expression of Gene Y in liver cells. They create three experimental conditions:
- Condition 1: Normal liver cells (TF-X present and functional)
- Condition 2: Liver cells with TF-X gene knocked out (no TF-X protein produced)
- Condition 3: Liver cells with a mutant TF-X that cannot bind to DNA
Gene Y mRNA levels were measured in all three conditions:
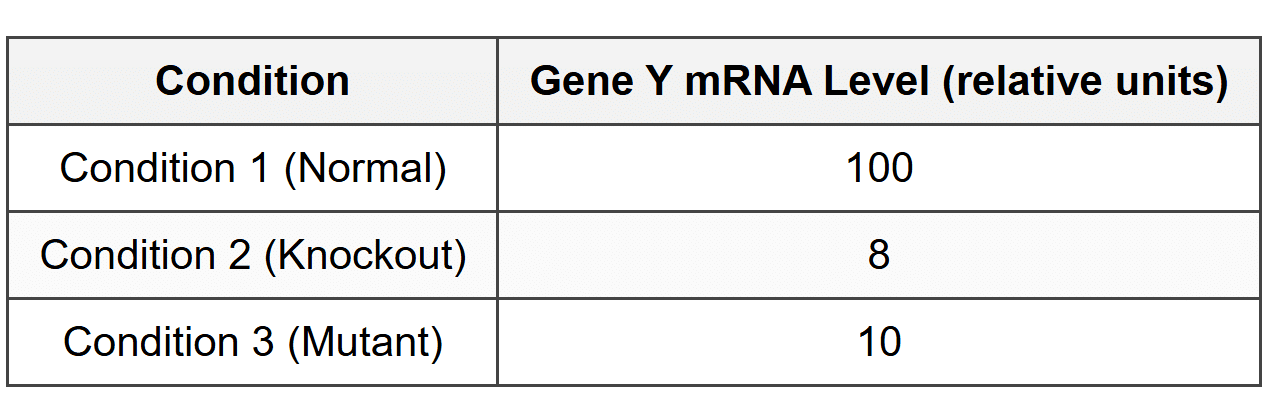
7. Based on these results, which conclusion about TF-X function is most strongly supported?
- TF-X functions as a repressor of Gene Y transcription.
- TF-X functions as an activator of Gene Y transcription and requires DNA binding for its function.
- TF-X has no significant effect on Gene Y expression in liver cells.
- TF-X regulates Gene Y expression exclusively through post-transcriptional mechanisms.
8. To further investigate the mechanism of TF-X action, researchers would most likely examine which of the following?
- Whether TF-X binds to specific DNA sequences in the promoter or enhancer regions of Gene Y.
- Whether TF-X affects the rate of DNA replication in liver cells.
- Whether TF-X is present in the ribosomal subunits during translation.
- Whether TF-X catalyzes the formation of peptide bonds between amino acids.
9. In eukaryotes, the TATA box is a DNA sequence found in many gene promoters, typically located about 25-30 base pairs upstream of the transcription start site. Which of the following best describes the function of the TATA box?
- It serves as the binding site for the ribosome to initiate translation.
- It provides a recognition and binding site for transcription factors and RNA polymerase II to initiate transcription.
- It marks the location where RNA splicing begins during pre-mRNA processing.
- It signals the termination of transcription and release of the mRNA transcript.
Questions 10-11 refer to the following data.
Figure 3: mRNA Stability and Gene Expression
Researchers studied two genes, Gene A and Gene B, which are transcribed at similar rates but produce different amounts of protein. They measured mRNA half-life and protein levels:
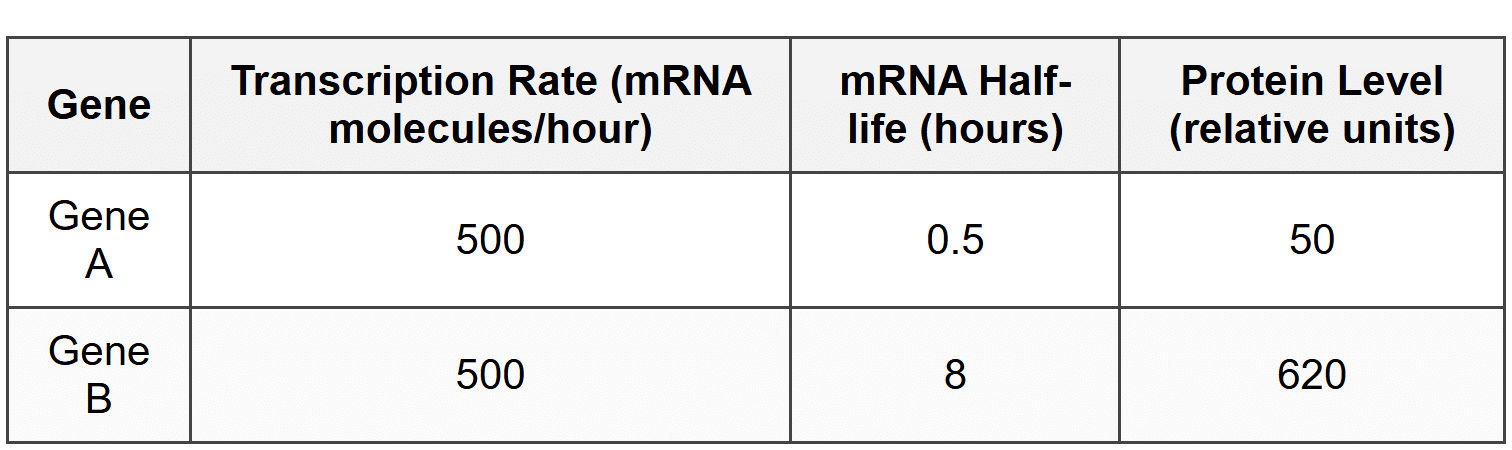
10. Which of the following best explains the difference in protein levels between Gene A and Gene B?
- Gene B mRNA is more stable and persists longer in the cell, allowing more time for translation to occur.
- Gene A has a higher transcription rate that saturates the translation machinery.
- Gene B produces proteins that are degraded more slowly than Gene A proteins.
- Gene A mRNA lacks a poly-A tail, which reduces its translation efficiency.
11. The poly-A tail at the 3' end of eukaryotic mRNA contributes to mRNA stability. A mutation that prevents poly-A tail addition to Gene B mRNA would most likely result in which of the following?
- An increase in Gene B protein levels due to faster translation.
- No change in Gene B protein levels because transcription rate remains constant.
- A decrease in Gene B protein levels due to more rapid mRNA degradation.
- Complete inability to transcribe Gene B because RNA polymerase cannot terminate transcription.
12. MicroRNAs (miRNAs) are small non-coding RNA molecules that regulate gene expression in eukaryotes. Which of the following mechanisms best describes how miRNAs typically function?
- miRNAs bind to complementary sequences on target mRNA molecules, leading to mRNA degradation or translational repression.
- miRNAs bind to DNA promoter regions and recruit RNA polymerase to increase transcription.
- miRNAs serve as templates for reverse transcriptase to produce complementary DNA strands.
- miRNAs replace damaged sections of mRNA through homologous recombination.
Questions 13-14 refer to the following scenario.
Epigenetic Modification Study
Researchers are investigating how DNA methylation affects gene expression. They compare two groups of cells:
- Group 1: Cells with normal methylation patterns
- Group 2: Cells treated with a drug that prevents DNA methylation
They measure the expression of Gene Z, which is normally methylated and silenced in Group 1 cells.
Results: Gene Z expression increased 15-fold in Group 2 cells compared to Group 1.
13. Which of the following best explains how DNA methylation regulates gene expression?
- Methylation of cytosine bases in promoter regions typically prevents transcription factor binding and gene expression.
- Methylation increases the rate of DNA replication by stabilizing the double helix structure.
- Methylation removes introns from pre-mRNA more efficiently during splicing.
- Methylation directly increases the binding affinity of RNA polymerase to promoters.
14. Which of the following scenarios would most likely involve changes in DNA methylation patterns?
- A point mutation in a gene coding sequence that changes a single amino acid.
- Differential gene expression during cellular differentiation from a stem cell to a specialized cell type.
- A deletion mutation that removes an entire gene from the chromosome.
- The formation of a peptide bond between two amino acids during translation.
15. In bacteria, several genes encoding enzymes for a single metabolic pathway are often organized into an operon, where they are transcribed as a single mRNA molecule. What is the primary advantage of this organization?
- It allows for more accurate DNA replication of these genes.
- It enables coordinated regulation of functionally related genes in response to environmental signals.
- It prevents mutations from occurring in these genes.
- It ensures that each gene produces a different amount of protein product.
Questions 16-17 refer to the following experiment.
Post-Transcriptional Regulation Experiment
Scientists studying iron metabolism identified an iron-responsive element (IRE) in the 5' untranslated region (UTR) of ferritin mRNA. Ferritin is a protein that stores iron in cells.
When cellular iron levels are low, an iron regulatory protein (IRP) binds to the IRE and blocks ribosome binding, preventing translation.
When cellular iron levels are high, iron binds to IRP, causing IRP to release from the IRE, allowing translation to proceed.
The graph below shows ferritin protein levels and ferritin mRNA levels under different iron concentrations:
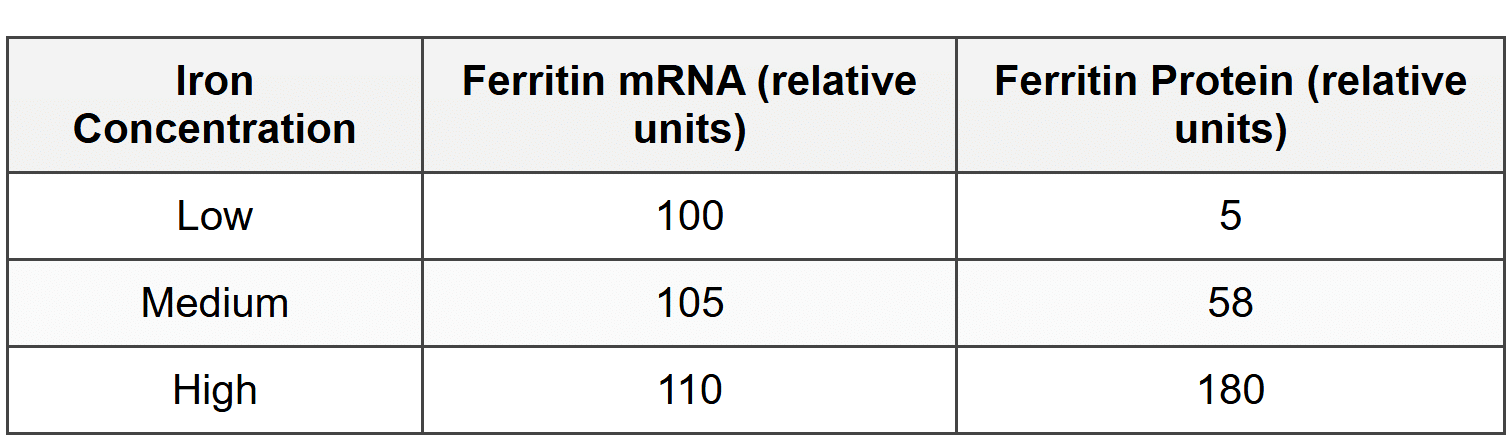
16. What level of gene regulation is primarily demonstrated in this experiment?
- Transcriptional regulation, because iron concentration affects mRNA production.
- Translational regulation, because ferritin protein levels change dramatically while mRNA levels remain relatively constant.
- Post-translational regulation, because iron directly modifies the ferritin protein structure.
- Epigenetic regulation, because iron causes methylation of the ferritin gene promoter.
17. This mechanism represents an example of which broader biological principle?
- Positive feedback, where increasing iron levels stimulate further iron uptake.
- Negative feedback, where accumulating ferritin protein inhibits its own production.
- Homeostatic regulation, where cells adjust protein production in response to changing conditions.
- Competitive inhibition, where iron competes with ferritin for binding sites on IRP.
18. Select TWO answers. Which TWO of the following are accurate differences between the regulation of gene expression in prokaryotes and eukaryotes?
- Prokaryotic gene regulation occurs primarily at the transcriptional level, while eukaryotic gene regulation occurs at multiple levels including transcriptional, post-transcriptional, translational, and post-translational.
- Eukaryotic genes can be regulated by chromatin remodeling and histone modifications, while prokaryotes lack histones and chromatin structure.
- Prokaryotic cells use transcription factors to regulate gene expression, while eukaryotic cells do not use transcription factors.
- Eukaryotic mRNA is translated while still being transcribed, while prokaryotic mRNA must be processed before translation.
Questions 19-20 refer to the following information.
Histone Modification and Gene Expression
Researchers investigated the effect of histone acetylation on gene expression. They measured the expression of Gene W under three conditions:
- Condition A: Normal cells (baseline acetylation)
- Condition B: Cells treated with a histone acetyltransferase (HAT) inhibitor, which reduces acetylation
- Condition C: Cells treated with a histone deacetylase (HDAC) inhibitor, which increases acetylation
Results:
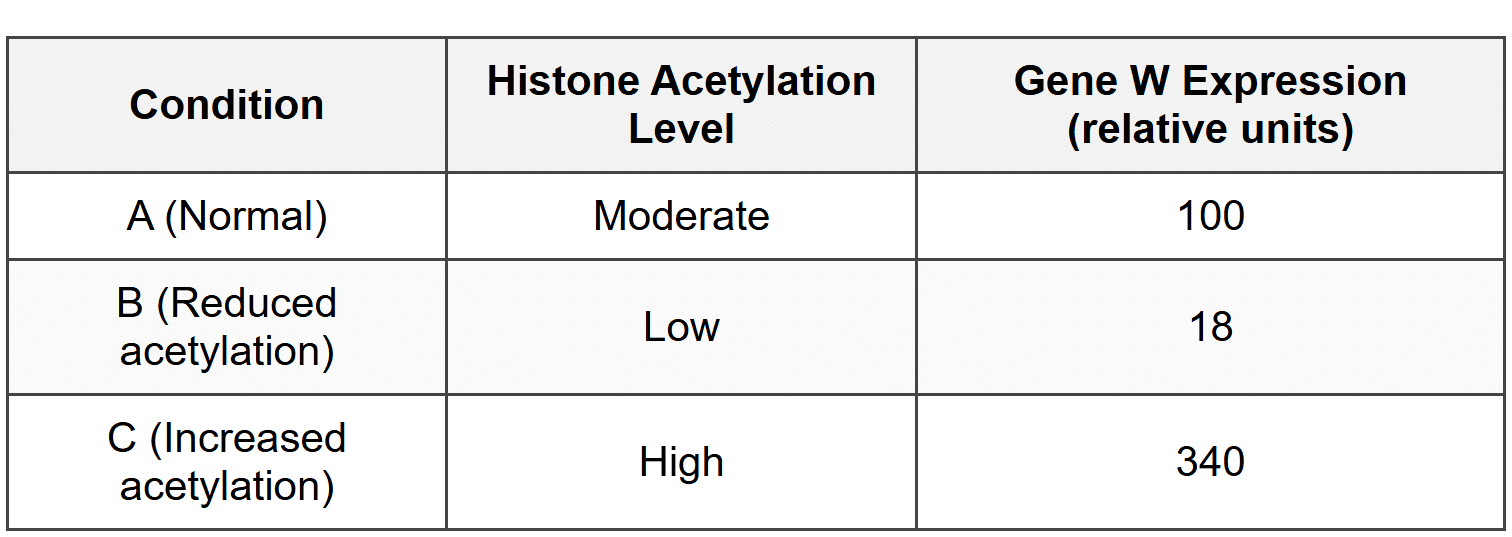
19. Based on these results, which of the following statements about histone acetylation is best supported?
- Histone acetylation decreases gene expression by condensing chromatin structure.
- Histone acetylation increases gene expression by loosening chromatin structure and making DNA more accessible to transcription factors.
- Histone acetylation has no significant effect on gene expression.
- Histone acetylation affects only the stability of mRNA, not the rate of transcription.
20. Acetyl groups added to histone proteins neutralize the positive charges on lysine residues. Which of the following best explains why this chemical change affects chromatin structure?
- Neutralizing positive charges weakens the electrostatic attraction between histones and the negatively charged DNA phosphate backbone, loosening the DNA-histone interaction.
- Neutralizing positive charges strengthens hydrogen bonding between adjacent nucleosomes, compacting the chromatin.
- Neutralizing positive charges causes DNA to change from B-form to Z-form, preventing transcription.
- Neutralizing positive charges activates DNA methylase enzymes that silence gene expression.
SECTION II: FREE RESPONSE
Directions
Answer both of the following questions. Where explanation or discussion is required, support your answers with relevant information and examples. Where calculation is required, show your work.
You have 40 minutes to answer both questions: approximately 22 minutes for Question 1 and approximately 18 minutes for Question 2.
Question 1 (Long FRQ - 10 points)
Background Information
The trp operon in E. coli encodes five enzymes involved in tryptophan biosynthesis. When tryptophan is abundant in the environment, the bacterium does not need to synthesize it. The operon is regulated by a repressor protein that is inactive on its own but becomes active when tryptophan (acting as a corepressor) binds to it.
A research team investigated the regulation of the trp operon by measuring enzyme activity under different conditions. They used three bacterial strains:
- Wild-type strain: Normal trp operon regulation
- Mutant strain 1: Mutation in the repressor gene that prevents tryptophan binding
- Mutant strain 2: Deletion of the operator sequence
Each strain was grown in two conditions: with tryptophan added to the medium (+Trp) or without tryptophan (-Trp). The activity of tryptophan biosynthesis enzymes was measured and results are shown below:
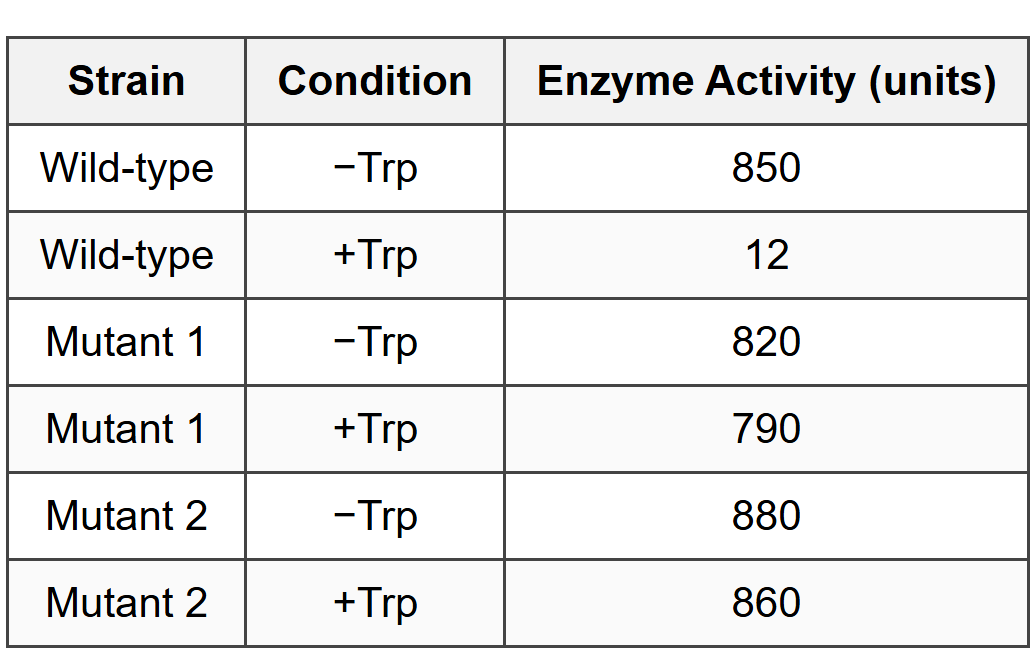
- Describe the mechanism by which the trp operon is regulated in wild-type E. coli when tryptophan is present in the environment. Your description should include the role of the repressor protein, the corepressor, and the operator sequence.
- Explain why enzyme activity remains high in Mutant strain 1 even when tryptophan is added to the medium, using the data to support your answer.
- Analyze the data for Mutant strain 2 and explain why deletion of the operator sequence results in the observed enzyme activity pattern.
- Calculate the fold-change in enzyme activity between the +Trp and -Trp conditions for the wild-type strain. Show your work.
- Design an experiment to determine whether the trp operon is also regulated at the level of mRNA stability (post-transcriptional regulation). Your experimental design should include:
- A description of the experimental procedure
- The measurements you would take
- How the results would support or refute the hypothesis that mRNA stability is involved in regulation
- Explain how the negative feedback regulation demonstrated by the trp operon represents an evolutionary advantage for bacteria living in environments where nutrient availability fluctuates.
Question 2 (Short FRQ - 4 points)
Gene expression can be regulated at multiple levels in eukaryotic cells, including transcriptional control, post-transcriptional control, translational control, and post-translational control.
- Identify one specific mechanism of post-transcriptional regulation in eukaryotes.
- Describe how this mechanism regulates gene expression, including the molecular components involved.
- Explain why having multiple levels of gene regulation provides an advantage to eukaryotic cells compared to regulating gene expression solely at the transcriptional level.
- Predict the effect on cellular function if a mutation disrupted the mechanism you described in part (b). Justify your prediction with specific reasoning.
ANSWER KEY
Part A - Multiple Choice Answers
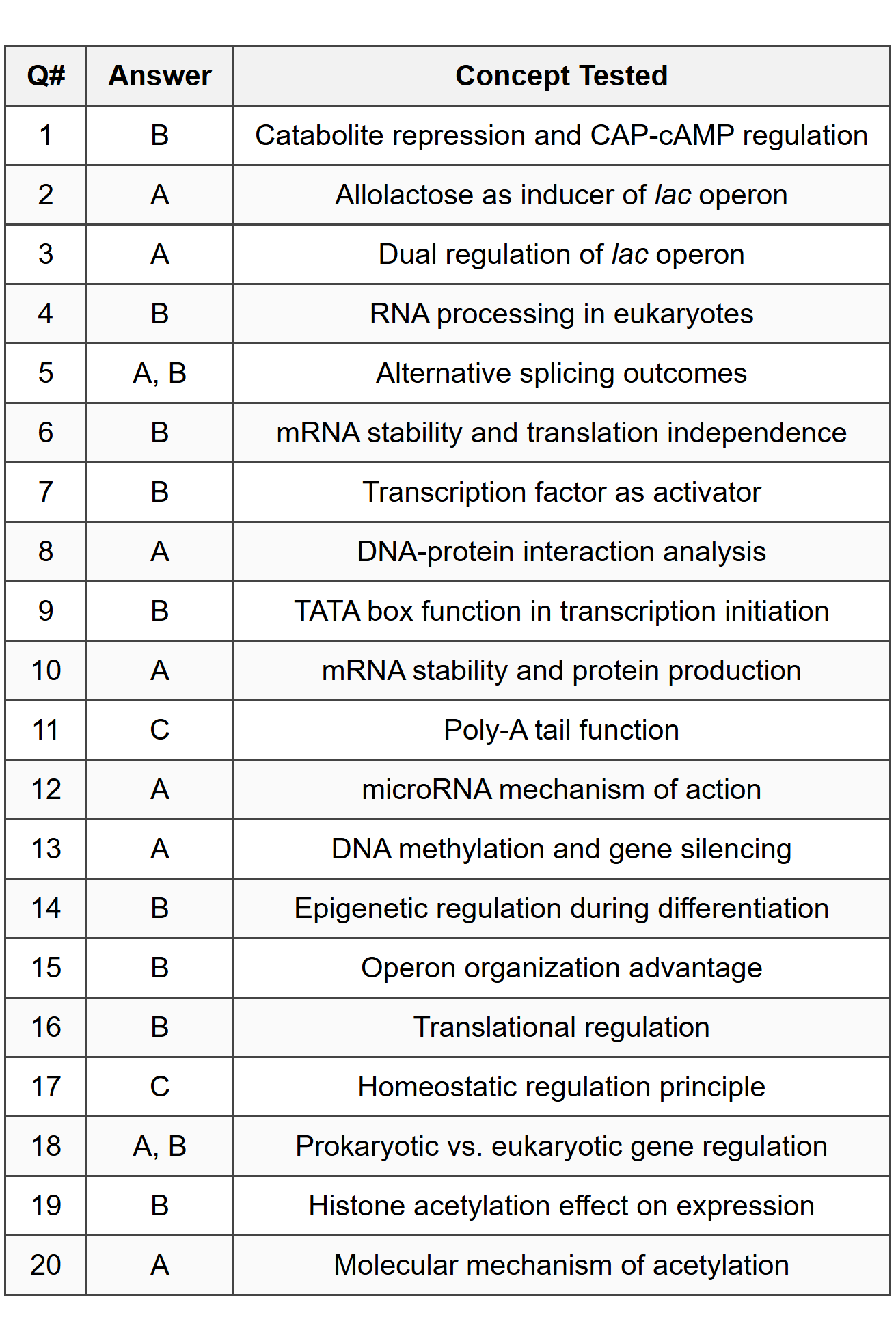
Part B - Free Response Answers
FRQ 1 - Answer Key
Part A: Mechanism of trp operon regulation (2 points)
Model Answer:
In wild-type E. coli, when tryptophan is present in the environment, it acts as a corepressor by binding to the inactive trp repressor protein. This binding causes a conformational change that allows the repressor-tryptophan complex to bind to the operator sequence located near the promoter of the trp operon. When the repressor is bound to the operator, it physically blocks RNA polymerase from transcribing the genes encoding tryptophan biosynthesis enzymes, effectively shutting down tryptophan production when it is not needed. This is an example of negative regulation and represents feedback inhibition at the transcriptional level.
Scoring note: Award 1 point for correctly describing the role of tryptophan as corepressor and the conformational change in the repressor. Award 1 point for explaining how repressor binding to the operator blocks transcription.
Part B: Explanation of Mutant strain 1 data (2 points)
Model Answer:
Mutant strain 1 has a mutation in the repressor gene that prevents tryptophan binding. The data show that enzyme activity remains high (790 units) even when tryptophan is added, compared to wild-type where enzyme activity drops to 12 units with tryptophan. This occurs because the mutant repressor cannot bind tryptophan as a corepressor, so it cannot undergo the conformational change needed to bind the operator sequence. Without functional repressor-operator binding, RNA polymerase can continuously transcribe the trp operon genes regardless of tryptophan availability. The cells waste energy synthesizing tryptophan biosynthesis enzymes even when tryptophan is abundant in the environment.
Scoring note: Award 1 point for connecting the mutation to inability of repressor to bind the operator. Award 1 point for explaining why this results in constitutive (constant) expression using specific data values.
Part C: Analysis of Mutant strain 2 (1 point)
Model Answer:
Mutant strain 2 lacks the operator sequence, which is the DNA binding site for the repressor protein. Without the operator, the repressor has no target site to bind to even when it is bound to tryptophan. The data show that enzyme activity is high in both -Trp (880 units) and +Trp (860 units) conditions, with minimal difference between them. This constitutive expression pattern demonstrates that the operator sequence is essential for repression to occur. The slight decrease from 880 to 860 units may reflect basal metabolic adjustments but indicates that transcriptional repression via the repressor-operator mechanism is completely lost.
Scoring note: Award 1 point for explaining that deletion of the operator prevents repressor binding and results in constitutive expression, supported by the data.
Part D: Calculate fold-change (1 point)
Model Answer:
Fold-change = (Enzyme activity in -Trp) ÷ (Enzyme activity in +Trp)
Fold-change = 850 units ÷ 12 units
Fold-change = 70.8-fold (or approximately 71-fold)
This means enzyme activity is about 71 times higher when tryptophan is absent compared to when it is present, demonstrating strong negative regulation by tryptophan.
Scoring note: Award 1 point for correct calculation with work shown. Accept answers from 70-71 fold. Units cancel out in this ratio.
Part E: Experimental design for mRNA stability (2 points)
Model Answer:
Procedure: Grow wild-type E. coli in medium without tryptophan to induce high trp operon transcription. At time zero, add rifampicin (an RNA polymerase inhibitor) to block all new transcription. Also at time zero, split the culture into two groups: one that receives added tryptophan (+Trp) and one that does not (-Trp control).
Measurements: Extract RNA samples at multiple time points (0, 5, 10, 15, 20, 30 minutes) and measure trp operon mRNA levels using quantitative RT-PCR or Northern blot analysis. Plot mRNA abundance over time for both conditions.
Interpretation: If mRNA stability is involved in regulation, the +Trp condition should show faster mRNA degradation (steeper decline) compared to the -Trp control. If both conditions show identical decay rates, this would indicate that tryptophan does not regulate mRNA stability, and regulation occurs only at the transcriptional level. A difference in mRNA half-life between conditions would support post-transcriptional regulation.
Scoring note: Award 1 point for a valid procedure that blocks transcription and allows measurement of existing mRNA decay. Award 1 point for explaining how results would distinguish between transcriptional-only versus post-transcriptional regulation.
Part F: Evolutionary advantage explanation (2 points)
Model Answer:
The negative feedback regulation of the trp operon provides a significant evolutionary advantage by allowing bacteria to conserve energy and resources in fluctuating environments. Synthesizing tryptophan requires five different enzymes and consumes ATP and biosynthetic precursors. When tryptophan is available in the environment (e.g., from degraded proteins), bacteria can import it rather than synthesizing it de novo. By shutting down the biosynthetic pathway when tryptophan is present, cells avoid the metabolic cost of producing unnecessary enzymes and intermediates.
In environments where nutrient availability changes frequently, bacteria with functional trp operon regulation can respond rapidly to changes-ramping up synthesis when tryptophan becomes scarce and shutting it down when it becomes abundant. This metabolic flexibility allows faster growth rates and better competition for limited resources compared to bacteria with constitutive expression that waste energy producing tryptophan unnecessarily. Natural selection would favor bacteria with efficient gene regulation, as they can allocate resources toward reproduction rather than unnecessary enzyme synthesis.
Scoring note: Award 1 point for explaining energy/resource conservation. Award 1 point for connecting regulation to competitive advantage and natural selection in variable environments.
FRQ 2 - Answer Key
Part A: Identify post-transcriptional mechanism (1 point)
Model Answer:
One specific mechanism of post-transcriptional regulation in eukaryotes is alternative splicing of pre-mRNA.
(Alternative acceptable answers: RNA splicing, 5' capping, 3' polyadenylation, mRNA localization, microRNA regulation, RNA-binding protein regulation, mRNA stability control)
Scoring note: Award 1 point for correctly identifying any valid post-transcriptional mechanism. Must be post-transcriptional, not transcriptional or translational.
Part B: Describe mechanism (1 point)
Model Answer (using alternative splicing):
Alternative splicing regulates gene expression by selectively including or excluding specific exons during pre-mRNA processing in the nucleus. The spliceosome, a complex of small nuclear ribonucleoproteins (snRNPs) and associated proteins, catalyzes the removal of introns and joining of exons. Different combinations of exons can be joined together in different cell types or developmental stages, controlled by splicing regulatory proteins such as SR proteins (serine/arginine-rich proteins) that promote exon inclusion and hnRNPs (heterogeneous nuclear ribonucleoproteins) that can promote exon skipping. This allows a single gene to produce multiple different mRNA variants, each potentially encoding a protein with different structure, function, or regulatory properties.
Scoring note: Award 1 point for describing the molecular mechanism, including the cellular machinery involved (spliceosome, regulatory proteins) and how it affects mRNA structure.
Part C: Explain advantage of multiple regulatory levels (1 point)
Model Answer:
Having multiple levels of gene regulation provides several advantages to eukaryotic cells. First, it allows for finer temporal control-transcriptional regulation is relatively slow (requiring transcription and processing), while translational or post-translational regulation can produce rapid responses to stimuli. Second, it provides greater specificity-different regulatory levels can respond to different signals, allowing integration of multiple inputs. Third, it increases efficiency-post-transcriptional mechanisms allow cells to modulate protein output from existing mRNA without the energy cost of new transcription. Finally, multiple regulatory checkpoints provide redundancy and precision, ensuring that gene expression is tightly controlled and can be fine-tuned in response to complex developmental or environmental cues.
Scoring note: Award 1 point for explaining at least two distinct advantages with clear reasoning about why multiple levels are beneficial compared to transcriptional control alone.
Part D: Predict effect of mutation and justify (1 point)
Model Answer (using alternative splicing):
If a mutation disrupted alternative splicing, cells would lose the ability to produce multiple protein variants from single genes. This could have severe consequences for cellular function, particularly in complex tissues like the nervous system where alternative splicing generates protein diversity essential for specialized functions. For example, if splicing regulatory proteins were mutated, certain exons might be constitutively included or excluded, producing only one mRNA variant instead of the normal repertoire. This could result in:
- Loss of tissue-specific protein isoforms needed for specialized cell functions
- Developmental abnormalities if different isoforms are required at different stages
- Reduced ability to respond to environmental changes that normally trigger splicing pattern changes
The severity would depend on which genes are affected, but broadly, the cell would lose an important mechanism for increasing proteomic diversity without expanding genome size, limiting its functional complexity.
Scoring note: Award 1 point for a specific, justified prediction that logically connects the loss of the mechanism to a cellular or organismal consequence. Must include reasoning, not just stating "it would be bad."





