NCERT Solutions: Biotechnology: Principles & Processes
Q1: Can you list 10 recombinant proteins which are used in medical practice? Find out where they are used as therapeutics (use the internet).
Ans: Recombinant proteins are produced using recombinant DNA technology, where a desired gene is inserted into an expression system to synthesise the protein in large amounts.
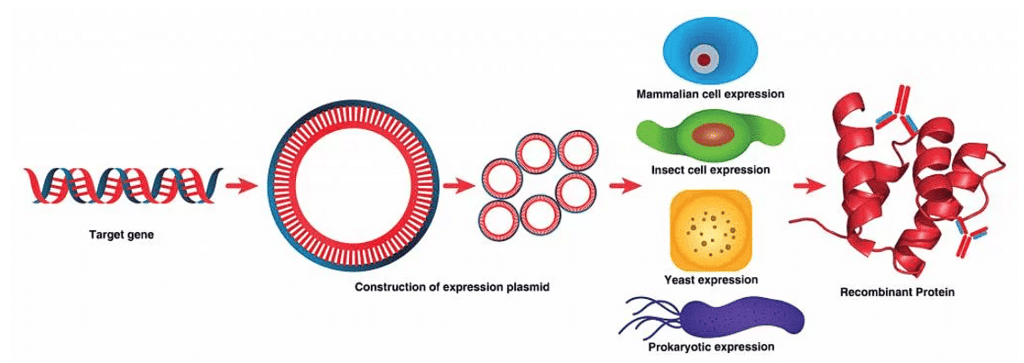 Recombinant Protein
Recombinant ProteinThis technology involves the transfer of specific genes from an organism into another organism using vectors and restriction enzymes as molecular tools.
Ten recombinant proteins used in medical practice are:
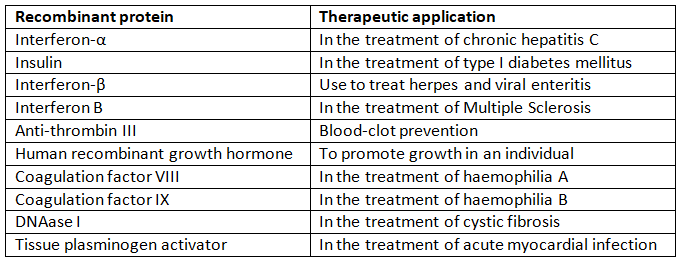
Q2: Make a chart (with diagrammatic representation) showing a restriction enzyme, the substrate DNA on which it acts, the site at which it cuts DNA and the product it produces.
Ans: The name of the restriction enzyme is BamHI.
Enhanced explanation: BamHI is a Type II restriction endonuclease that recognises the palindromic sequence 5′-GGATCC-3′ in double-stranded DNA. It cleaves between the two G nucleotides on each strand to produce fragments with 5′ overhangs (sticky ends).
Substrate DNA: Double-stranded DNA containing the sequence 5′-GGATCC-3′.
Recognition site: 5′-G G A T C C-3′ (palindromic).
Cut position: Between the first and second G on each strand, producing a 4-base 5′ overhang: 5′-G A T C-3′.
Product: Two DNA fragments with complementary sticky ends (5′ overhangs) that facilitate ligation with complementary sequences.
Q3: From what you have learnt, can you tell whether enzymes are bigger or DNA is bigger in molecular size? How did you know?
Ans: DNA molecules are generally much larger than individual enzyme molecules.
Reasoning:
• A single DNA molecule (for example, a chromosome) may contain millions to hundreds of millions of nucleotide base pairs, giving it a very large molecular mass.
• Most enzymes are single proteins of a few hundred to a few thousand amino acids; their molecular masses are much smaller compared with a whole chromosome or a full-length genomic DNA molecule.
• For example, a human chromosome contains millions of base pairs while a typical enzyme contains a few hundred amino acids. Therefore, in molecular size and mass, DNA (chromosomal molecules) is larger than enzymes.
Q 4: What would be the molar concentration of human DNA in a human cell? Consult your teacher.
Ans: The molar concentration of human DNA in a human diploid cell is as follows:
⇒ Total number of chromosomes × 6.023 × 1023
⇒ 46 × 6.023 × 1023
⇒ 2.77 ×1018 Moles
Hence, the molar concentration of DNA in each diploid cell in humans is 2.77 × 10 23 moles.
Q5: Do eukaryotic cells have restriction endonucleases? Justify your answer.
Ans: Classically, the well-characterised restriction-modification systems (restriction endonucleases paired with methylases) occur in prokaryotes, especially bacteria. These enzymes protect bacteria against invading viral (phage) DNA by recognising and cutting specific unmethylated sequences.
Justification:
• Restriction endonucleases used in molecular biology were originally isolated from bacteria and form part of bacterial defence systems.
• Eukaryotic genomes do have nucleases (enzymes that cut nucleic acids) and DNA-modifying enzymes, and eukaryotic DNA is often methylated, which alters recognition by some nucleases.
• However, eukaryotes do not possess the classical restriction-methylation systems that protect against phage infection; therefore, in the usual sense of restriction endonucleases used in cloning, these are characteristic of prokaryotes.
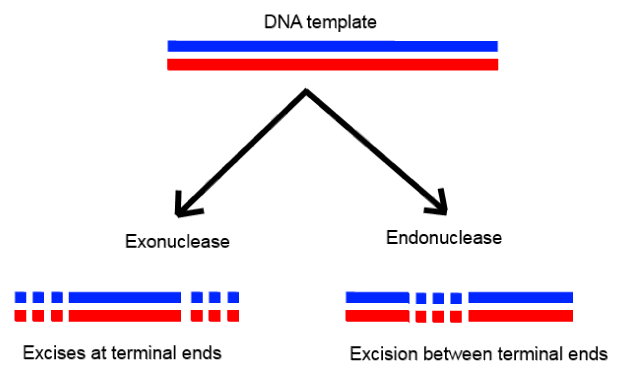 Endonucleases
EndonucleasesQ6: Besides better aeration and mixing properties, what other advantages do stirred tank bioreactors have over shake flasks?
Ans: The shake flask method is used for small-scale culture in the laboratory, while stirred-tank bioreactors are designed for large-scale production. In addition to improved aeration and mixing, stirred-tank bioreactors offer several other advantages:
(1) Controlled sampling: Small volumes of culture can be withdrawn through a sterile sampling port for testing without contaminating the whole culture.
(2) Foam control: They are fitted with foam breakers or antifoam systems to manage foam formation.
(3) Environmental control: Integrated systems regulate and maintain temperature, pH and dissolved oxygen precisely.
(4) Scale-up and reproducibility: Conditions can be scaled reliably from laboratory to industrial volumes, giving consistent product yields.
(5) Sterility and monitoring: Stirred reactors are closed systems with better sterility, online sensors and automated control for long culture runs.
(6) Better control of shear and mixing profiles depending on impeller design, which helps sensitive cells or higher-density cultures.
 Stirred Tank Bioreactors
Stirred Tank Bioreactors
Q7: Collect 5 examples of palindromic DNA sequences by consulting your teacher. Better try to create a palindromic sequence by following base-pair rules.
Ans: A palindromic DNA sequence reads the same on complementary strands when both are read in the 5′→3′ direction. Such sites are commonly recognised by restriction enzymes.
Five examples of palindromic sequences are :
Q8: Can you recall meiosis and indicate at what stage a recombinant DNA is made?
Ans: Recombination (formation of recombinant DNA via crossing over) occurs during prophase I of meiosis, specifically at the pachytene stage.
Explanation:
• During pachytene, homologous chromosomes are fully synapsed and exchange corresponding segments between non-sister chromatids at points called chiasmata.
• This exchange mixes parental alleles and produces recombinant chromosomes, increasing genetic variation in gametes.
Q9: Can you think and answer how a reporter enzyme can be used to monitor transformation of host cells by foreign DNA in addition to a selectable marker?
Ans: A reporter gene is included in a DNA construct to indicate whether the foreign gene has been taken up and/or is being expressed by the host cell. Reporter genes give an easy, visible or measurable readout.
How reporters are used:
The reporter gene is placed on the same construct as the gene of interest. After transformation, cells that express the reporter indicate successful uptake and expression of the construct.
Examples of reporter genes and their use:
- lacZ encodes β-galactosidase; blue/white screening on X-gal plates distinguishes colonies with functional lacZ (blue) from those lacking it (white).
- GFP (green fluorescent protein) from Aequorea victoria emits green fluorescence under UV/blue light and indicates expression in live cells.
- Luciferase produces bioluminescence measurable with a luminometer, useful for sensitive assays.
Difference from selectable markers:
Selectable markers (e.g., antibiotic resistance) enable selection of cells that have taken up the construct, while reporter genes provide direct evidence of expression and can give quantitative or visual readouts of promoter activity or localisation.
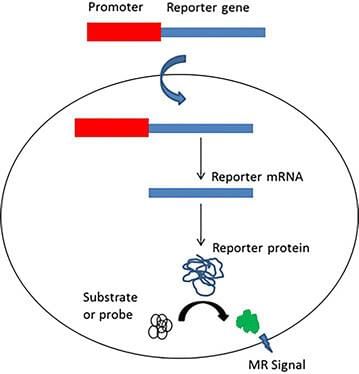 Reporter Gene
Reporter GeneThe researchers place the reporter gene and the foreign gene in the same DNA construct. Then, this combined DNA construct is inserted in the cell.
Here, the reporter gene is used as a selectable marker to find out the successful uptake of genes of interest (foreign genes). An example of reporter genes includes lac Z gene, which encodes a green fluorescent protein in a jelly fish.
Q10: Describe briefly the followings:
(a) Origin of replication
(b) Bioreactors
(c) Downstream processing
Ans: (a) Origin of replication: Origin of replication (ori) is a specific DNA sequence where the process of DNA replication begins. Proteins that initiate replication recognise this site, unwind the double helix and recruit DNA polymerases. Replication from an origin may proceed uni-directionally or bi-directionally depending on the organism and origin type.
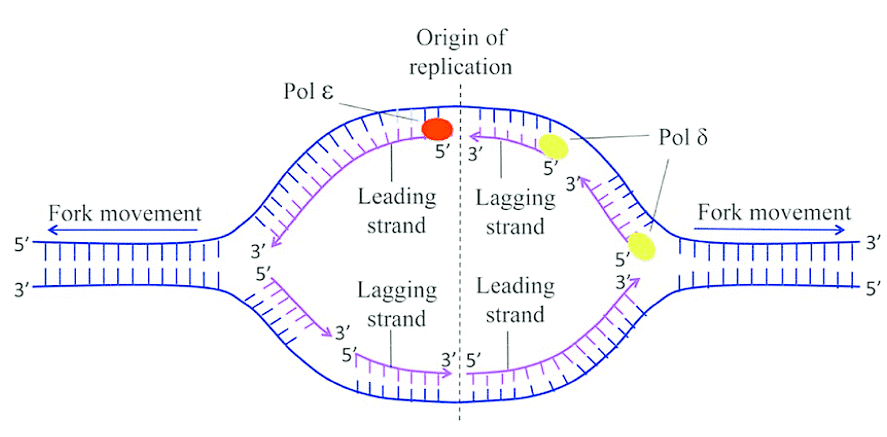 Origin of Replication
Origin of Replication(b) Bioreactors: Bioreactors are engineered vessels used for controlled, large-scale cultivation of cells or microorganisms to produce desired products (enzymes, proteins, vaccines, metabolites). They maintain optimal environmental conditions (temperature, pH, dissolved oxygen, nutrient supply) and include components such as agitation systems, aeration systems, sensors, foam control and sampling ports.
(c) Downstream processing: Downstream processing refers to the sequence of operations used to recover and purify the product after biosynthesis. Typical steps include cell removal (centrifugation or filtration), product release (if intracellular), capture and initial concentration, purification (chromatography, precipitation), polishing steps to achieve required purity, and final formulation and sterilisation for clinical or commercial use. Quality checks and validation are done throughout the process.
Q11: Explain briefly
(a) PCR
(b) Restriction enzymes and DNA
(c) Chitinase
Ans: (a) PCR: Polymerase chain reaction (PCR) is an in-vitro method to amplify a specific DNA segment into millions of copies. The main steps in a PCR cycle are:
• Denaturation - Heat the double-stranded DNA to separate the strands.
• Annealing - Cool the reaction so primers bind (anneal) to complementary sequences on each single strand.
• Extension - A thermostable DNA polymerase (e.g., Taq polymerase from Thermus aquaticus) extends the primers by adding dNTPs to synthesise new DNA strands.
Repeating these cycles (typically 20-35 cycles) results in exponential amplification of the target fragment.
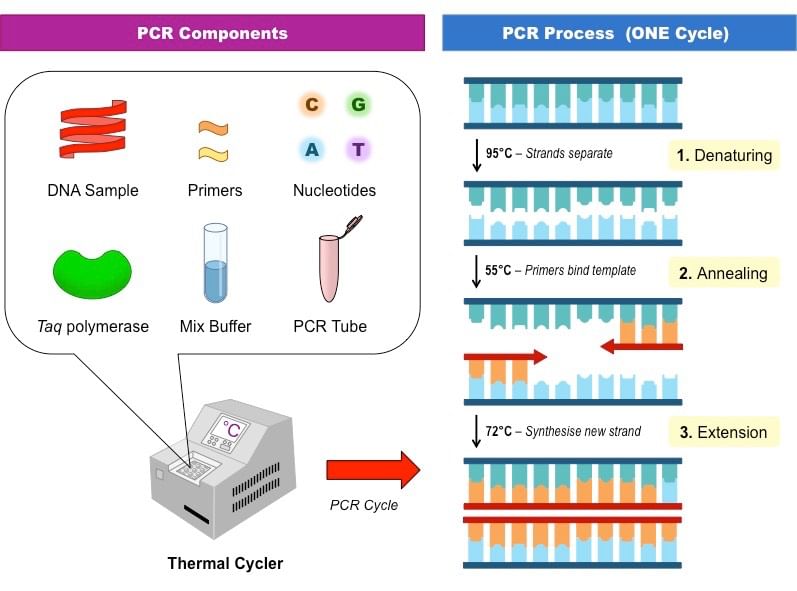 Polymerase chain reaction
Polymerase chain reactionThe process involves in-vitro synthesis of sequences using a primer, a template strand, and a thermo stable DNA polymerase enzyme obtained from a bacterium, called Thermus aquaticus. The enzyme utilises building blocks dNTPs (deoxynucleotides) to extend the primer.
In the first step, the double stranded DNA molecules are heated to a high temperature so that the two strands separate into a single stranded DNA molecule. This process is called denaturation.
Then, this ssDNA molecule is used as a template strand for the synthesis of a new strand by the DNA polymerase enzyme and this process is called annealing, which results in the duplication of the original DNA molecule.
This process is repeated over several cycles to obtain multiple copies of the rDNA fragment.
(b) Restriction enzymes are molecular scissors that recognise specific short DNA sequences (recognition sites) and cut the DNA at or near those sites. They are essential tools in genetic engineering because they generate predictable fragments with either blunt or sticky (overhanging) ends. For example, EcoRI recognises 5′-GAATTC-3′ and cuts between G and A to generate 5′ overhangs. These fragments can be joined using DNA ligase to form recombinant DNA molecules.
Types: Exonucleases remove nucleotides from the ends of DNA, while endonucleases cut within the DNA strand.
(c) Chitinase: Chitinase is an enzyme that degrades chitin, a polymer found in fungal cell walls and arthropod exoskeletons. In biotechnology, chitinase can be used to break down fungal cell walls to release intracellular contents (for example, during DNA isolation from fungi) or as a biocontrol agent against fungal pathogens.
Q12: Discuss with your teacher and find out how to distinguish between
(a) Plasmid DNA and chromosomal DNA
(b) RNA and DNA
(c) Exonuclease and Endonuclease
Ans: (a) Plasmid DNA and chromosomal DNA
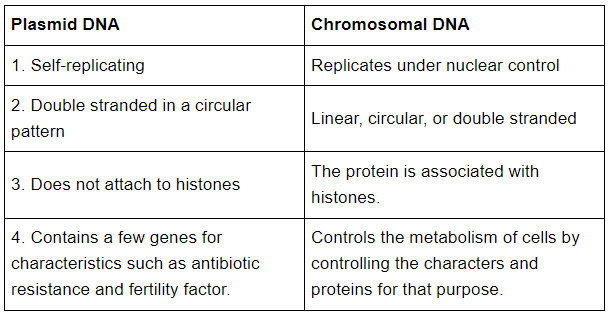
(b) RNA and DNA
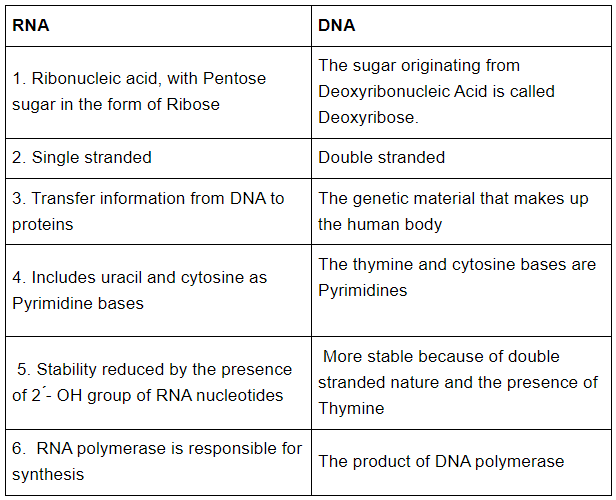
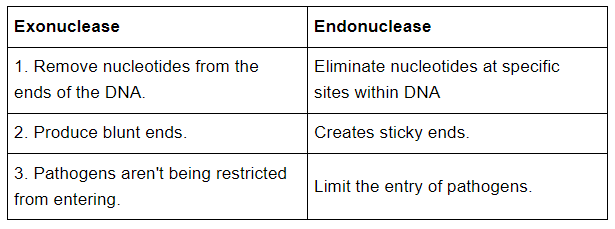
(c) Exonuclease and Endonuclease
FAQs on NCERT Solutions: Biotechnology: Principles & Processes
| 1. What are the basic principles of biotechnology? |  |
| 2. How are recombinant DNA technologies used in biotechnology processes? |  |
| 3. What is the significance of bioprocessing in biotechnology? |  |
| 4. How are enzymes utilized in biotechnology processes? |  |
| 5. How does biotechnology contribute to sustainable agriculture practices? |  |






