Protein and Structure of Protein | Biology Class 11 - NEET PDF Download
| Table of contents |

|
| What is Protein? |

|
| Structure of Protein |

|
| (a) Primary Structure |

|
| (b) Secondary Structure |

|
| (c) Tertiary Structure |

|
| (d) Quaternary Structure |

|
| Functions of Proteins |

|
What is Protein?
Proteins are large, complex molecules made up of chains of smaller units called amino acids. They are essential for the structure, function, and regulation of the body's cells, tissues, and organs.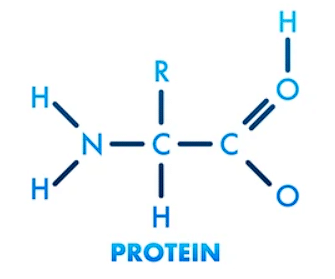
- Proteins are essentially chains of amino acids linked together by peptide bonds. These chains, called polypeptides, are made up of various combinations of 20 different types of amino acids.
- Because of this diversity in amino acid composition, proteins are referred to as heteropolymers rather than homopolymers, which consist of repeating units of a single type of monomer.
- Understanding the types of amino acids in proteins is crucial, especially when it comes to nutrition. Some amino acids are deemed essential because our bodies cannot produce them, so we must obtain them from our diet. Proteins derived from food serve as the source of these essential amino acids. In contrast, non-essential amino acids are those our bodies can synthesize on their own.
- Proteins play numerous vital roles in living organisms, such as facilitating nutrient transport across cell membranes, combating infectious agents, acting as hormones, and serving as catalysts for biochemical reactions as enzymes.
Structure of Protein
Proteins are biopolymers consisting of a large number of amino acids linked together through peptide bonds having three-dimensional (3D) structures. The structure of proteins is very complex. Protein structure and shape can generally be studied at four different levels, namely primary, secondary, tertiary, and quaternary structures.
(a) Primary Structure
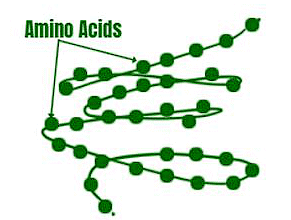
Think of a protein as a line of beads where each bead represents an amino acid. The order of these beads, from beginning to end, is the primary structure. The first bead is called the N-terminal amino acid, and the last one is the C-terminal amino acid.
(b) Secondary Structure
Instead of being a straight line, the protein folds and twists into shapes like a helix (think of a spiral staircase) or other structures. This folding is the secondary structure.
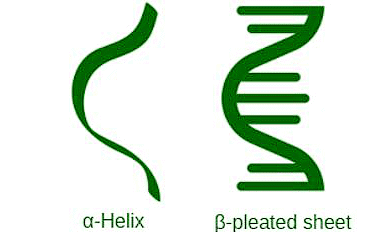
- α – Helix structure: The ∝-helix model was proposed by Linus Pauling in 1951 based on theoretical considerations. However, was later verified experimentally. It is the majority generic form in which a polypeptide chain forms all possible types of hydrogen bonds by twisting into a right-handed screw helix) with the -NH group of each amino acid residue hydrogen-bonded to the C=0 group of an imminent turn of the helix. This is called ∝-helix. This structure can be conjectured as if one can coil a polypeptide chain next to an invisible cylinder.
- β-pleated sheet structure: In this structure, all polypeptide chains are stretched out to almost supreme extension and then lay down side by side in a zig-zag manner to form a flat sheet. Each sheet is held to bonds. These sheets are stacked in one structure called β-pleated sheet structure. The structure resembles the pleated stratum of the drapery and, as closely as possible, is referred to as a β-pleated sheet.
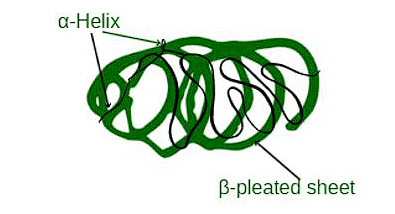
(c) Tertiary Structure
Now imagine taking that twisted line and folding it up into a 3D shape, like a crumpled ball of yarn. This final shape is the tertiary structure, and it's crucial for the protein to do its job.
(d) Quaternary Structure
Sometimes, proteins are made up of more than one twisted line. The arrangement of these lines with each other forms the quaternary structure. For example, adult human hemoglobin has four parts, two of which are the same and two are different. These parts fit together like puzzle pieces to form the hemoglobin protein.
Functions of Proteins
- They help in the maintenance and growth of tissues.
- The proteins known as enzymes, help in performing many biochemical reactions within the external of the cells.
- The hormones are the proteins, also known as the chemical messengers, which help in communication between the tissues, cells, and organs.
- Some proteins are fibrous, which provides stiffness and rigidity to the cells and tissues.
- They perform an essential function in monitoring the concentrations of bases and acids in the blood and other fluids found in the body.
- Globulin and albumin are the proteins found in the blood, which assist in maintaining the balance of the body fluids.
|
150 videos|398 docs|136 tests
|
FAQs on Protein and Structure of Protein - Biology Class 11 - NEET
| 1. What are the four levels of protein structure? |  |
| 2. What are the functions of proteins in the body? |  |
| 3. How are amino acids classified? |  |
| 4. What is the configuration of a protein molecule? |  |
| 5. Why is understanding protein structure important in biology and medicine? |  |
















