31 Years NEET Previous Year Questions: Biotechnology: Principles & Processes - Grade 12 MCQ
25 Questions MCQ Test - 31 Years NEET Previous Year Questions: Biotechnology: Principles & Processes
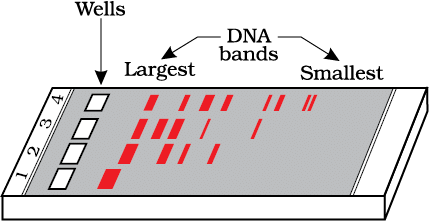
DNA strands on a gel stained with ethidium bromide when viewed under UV radiation, appear as: [NEET 2021]
Genes of interest can be selected from a genomic library by using
[NEET Kar. 2013]
| 1 Crore+ students have signed up on EduRev. Have you? Download the App |
The colonies of recombinant bacteria appear white in contrast to blue colonies of non recombinant bacteria because of :
[NEET 2013]
DNA fragments generated by the restriction endonucleases in a chemical reaction can be separated by :
[NEET 2013]
Which one of the following represents a palindromic sequence in DNA?
[2012M]
The figure below shows three steps (A, B, C) of Polymerase Chain Reaction (PCR). Select the option giving correct identification together with what it represents?
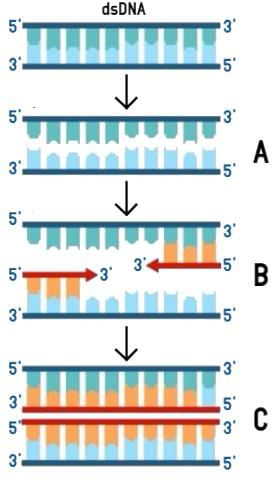
[2012M]
In genetic engineering, the antibiotics are used
[2012M]
Biolistics (gene-gun) is suitable for
[2012M]
For transformation, micro-particles coated with DNA to be bombarded with gene gun are made up of :
[2012]
Which one is a true statement regarding DNA polymerase used in PCR
[2012]
A single strand of nucleic acid tagged with a radioactive molecule is called :
[2012]
PCR and Restriction Fragment Length Polymorphism are the methods for :
[2012]
The figure below is the diagrammatic representation of the E.Coli vector pBR 322. Which one of the given options correctly identifies its certain component (s) ?
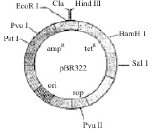
[2012]
Bacillus thuringiensis forms protein crystals which contain insecticidal protein.
[2011M]
Agarose extracted from sea weeds finds use in :
[2011]
There is a restriction endonuclease called EcoRI. What does .co. part in it stand for ?
[2011]
Restriction endonucleases are enzymes which
[2010]
DNA or RNA segment tagged with a radioactive molecule is called
[2010]
Which one of the following palindromic base sequences in DNA can be easily cut at about the middle by some particular restriction enzyme?
[2010]
Which one of the following is used as vector for cloning genes into higher organisms?
[2010]
Select the correct statement from the following?
[2010]
Polyethylene glycol method is used for
[2009]
The linking of antibiotic resistance gene with the plasmid vector became possible with
[2008]
Gel electrophoresis is used for
Introduction of food plants developed by genetic engineering is not desirable because
[2002]