All Exams > NEET > 30-Day Revision Course for NEET > All Questions
All questions of Biotechnology and Its Processes for NEET Exam
Study the following statements regarding recombinant DNA technology and select the incorrect ones
(i) Taq polymerase extends the primers using the nucleotides provided in the reaction
(ii) Antibiotic resistance genes are considered as desirable genes in recombinant DNA technology
(iii) DNA fragments are separated according to their charge only, in agarose gel electrophoresis
(iv) Transformation is a procedure through which a piece of DNA is integrated into the genome of a host bacterium
(v) To produce higher yields of the desired protein, host cells can be multiplied in a continuous culture
(vi) Downstream processing is one of the steps of polymerase chain reaction- a)(i), (iii) and (vi)
- b)(i), (iii) and (v)
- c)(ii), (iii) and (v)
- d)(i), (iv) and (v)
Correct answer is option 'A'. Can you explain this answer?
Study the following statements regarding recombinant DNA technology and select the incorrect ones
(i) Taq polymerase extends the primers using the nucleotides provided in the reaction
(ii) Antibiotic resistance genes are considered as desirable genes in recombinant DNA technology
(iii) DNA fragments are separated according to their charge only, in agarose gel electrophoresis
(iv) Transformation is a procedure through which a piece of DNA is integrated into the genome of a host bacterium
(v) To produce higher yields of the desired protein, host cells can be multiplied in a continuous culture
(vi) Downstream processing is one of the steps of polymerase chain reaction
(i) Taq polymerase extends the primers using the nucleotides provided in the reaction
(ii) Antibiotic resistance genes are considered as desirable genes in recombinant DNA technology
(iii) DNA fragments are separated according to their charge only, in agarose gel electrophoresis
(iv) Transformation is a procedure through which a piece of DNA is integrated into the genome of a host bacterium
(v) To produce higher yields of the desired protein, host cells can be multiplied in a continuous culture
(vi) Downstream processing is one of the steps of polymerase chain reaction
a)
(i), (iii) and (vi)
b)
(i), (iii) and (v)
c)
(ii), (iii) and (v)
d)
(i), (iv) and (v)
| | Mira Joshi answered |
Antibiotic resistance genes are selectable markers. Desirable genes are the ones which are introduced in the vector for getting desired protein product. In agarose gel electrophoresis, DNA fragments are separated according to their charge and size. After the formation of the product in a bioreactors, it undergoes through some processes before a finished product is ready for marketing. The processes include separation and purification of products which are collectively called as downstream processing.
After completion of the biosynthetic stage in the bioreactors, the product undergoes separation and purification processes, collectively termed as __________.- a)Transformation
- b)Electrophoresis
- c)Downstream processing
- d)Upstream processing
Correct answer is option 'C'. Can you explain this answer?
After completion of the biosynthetic stage in the bioreactors, the product undergoes separation and purification processes, collectively termed as __________.
a)
Transformation
b)
Electrophoresis
c)
Downstream processing
d)
Upstream processing
| | Dev Patel answered |
After the formation of the product in bioreactors, it undergoes through some processes before a finished product is ready for marketing. The processes include separation and purification of products which is collectively called as downstream processing.
Plasmid used to construct the first recombinant DNA was isolated from which bacterium species?- a)Escherichia coli
- b)Salmonella typhimurium
- c)Agrobacterium tumefaciens
- d)Thermus aquaticus
Correct answer is option 'B'. Can you explain this answer?
Plasmid used to construct the first recombinant DNA was isolated from which bacterium species?
a)
Escherichia coli
b)
Salmonella typhimurium
c)
Agrobacterium tumefaciens
d)
Thermus aquaticus
| | Riya Banerjee answered |
The first recombinant DNA was constructed by Stanley Cohen and Herbert Boyer in 1972. They cut the piece of DNA from a plasmid carrying antibiotic resistance gene in the bacterium Salmonella typhimurium and linked it to the plasmid of Escherichia coil.
Read statements (i)-(iv). Which of the following statements are incorrect?
(i) First transgenic buffalo Rosie produced milk which was human alpha-lactalbumin enriched.
(ii) Restriction enzymes are used in isolation of DNA from other macromolecules.
(iii) Downstream processing is one of the steps of rDNA technology.
(iv) Disarmed pathogen vectors are also used in the transfer of rDNA into the host.- a)(ii) and (iii)
- b)(iii) and (iv)
- c)(i) and (iii)
- d)(i) and (ii)
Correct answer is option 'D'. Can you explain this answer?
Read statements (i)-(iv). Which of the following statements are incorrect?
(i) First transgenic buffalo Rosie produced milk which was human alpha-lactalbumin enriched.
(ii) Restriction enzymes are used in isolation of DNA from other macromolecules.
(iii) Downstream processing is one of the steps of rDNA technology.
(iv) Disarmed pathogen vectors are also used in the transfer of rDNA into the host.
(i) First transgenic buffalo Rosie produced milk which was human alpha-lactalbumin enriched.
(ii) Restriction enzymes are used in isolation of DNA from other macromolecules.
(iii) Downstream processing is one of the steps of rDNA technology.
(iv) Disarmed pathogen vectors are also used in the transfer of rDNA into the host.
a)
(ii) and (iii)
b)
(iii) and (iv)
c)
(i) and (iii)
d)
(i) and (ii)
| | Hansa Sharma answered |
In 1997, Rosie, the tirst transgenic cow was engineered to produce milk enriched with a human protein called alpha-lactalbumin, making it nutritionally more balanced. Restriction enzymes are used to cut DNA at specific sites.
The term 'recombinant DNA' refers to- a)DNA of the host cell
- b)DNA with a piece of foreign DNA
- c)DNA with selectable marker
- d)DNA with more than one recognition sites
Correct answer is option 'B'. Can you explain this answer?
The term 'recombinant DNA' refers to
a)
DNA of the host cell
b)
DNA with a piece of foreign DNA
c)
DNA with selectable marker
d)
DNA with more than one recognition sites
| | Rajat Parihar answered |
Construction of recombinant DNA, in which a foreign DNA fragment is inserted into a plasmid vector.
One of the key factors, which makes the plasmid the vector in genetic engineering is- a)Its resistance to antibiotics
- b)Its resistance to restriction enzymes
- c)Its ability to carry a foreign gene
- d)Its ability to cause infection in the host
Correct answer is option 'C'. Can you explain this answer?
One of the key factors, which makes the plasmid the vector in genetic engineering is
a)
Its resistance to antibiotics
b)
Its resistance to restriction enzymes
c)
Its ability to carry a foreign gene
d)
Its ability to cause infection in the host
| | Ajay Yadav answered |
Plasmids are extra-chromosomal, self-replicating, usually circular, double-stranded DNA molecules found naturally in many bacteria and also in some yeasts. Plasmids are usually not essential for normal cell growth and division, they often confer some traits to the host organism e.g., resistance to certain antibiotics. The plasmid that is used as a carrier for transferring a fragment of foreign DNA into a suitable host is called vehicle DNA or cloning vector or gene carrier.
Which of the following statement is not correct?- a)Recombinant technologies are used to produce desirable proteins
- b)Agrobacterium is a genus of bacteria that causes tumours in plants
- c)Log phase does not show any significant increase in the number of cells whereas the lag phase shows rapid multiplication of cells
- d)Dolly, a sheep was the first animal to be cloned in 1997
Correct answer is option 'C'. Can you explain this answer?
Which of the following statement is not correct?
a)
Recombinant technologies are used to produce desirable proteins
b)
Agrobacterium is a genus of bacteria that causes tumours in plants
c)
Log phase does not show any significant increase in the number of cells whereas the lag phase shows rapid multiplication of cells
d)
Dolly, a sheep was the first animal to be cloned in 1997
| | Ananya Das answered |
The statement that is not correct is:
3. Log phase does not show any significant increase in the number of cells whereas the lag phase shows rapid multiplication of cells.
Here's why:
- Log phase (or exponential phase) is actually characterized by a rapid and significant increase in the number of cells. During this phase, cells are dividing at an exponential rate.
- Lag phase is the initial phase where cells are adapting to new conditions and preparing for growth. During this phase, there is little to no significant increase in cell number as the cells are not yet dividing rapidly.
To summarize, the log phase is when cell multiplication is most rapid, not the lag phase.
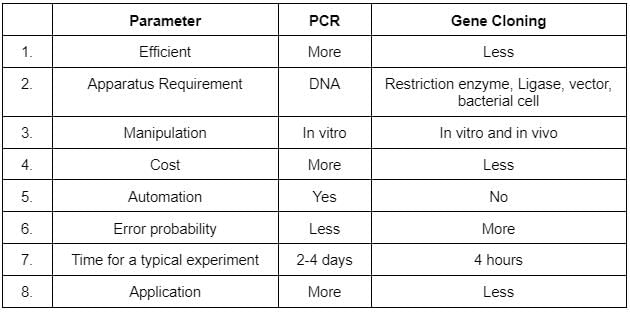
Given table gives an account of differences between PCR and gene cloning.. Which of the following points show the incorrect differences?
- a)1 and 3
- b)4 and 6
- c)4 and 7
- d)4, 7 and 8
Correct answer is option 'C'. Can you explain this answer?
Given table gives an account of differences between PCR and gene cloning.. Which of the following points show the incorrect differences?


a)
1 and 3
b)
4 and 6
c)
4 and 7
d)
4, 7 and 8
 | EduRev NEET answered |
The correct option is C 4 and 7
The cost of gene cloning is far more than PCR because gene cloning requires many intricate steps. PCR takes less than 4 hours while gene cloning can take days.
The cost of gene cloning is far more than PCR because gene cloning requires many intricate steps. PCR takes less than 4 hours while gene cloning can take days.
Which of the following are the types of bioreactors?
(i) Simple stirred-tank bioreactor
(ii) Complex Stirred-tank bioreactor
(iii) Sparged stirred-tank bioreactor
(iv) Agitator stirred-tank bioreactor- a)(i) and (iii)
- b)Only (iii)
- c)(i) and (ii)
- d)(i) and (ii) and (iv)
Correct answer is option 'A'. Can you explain this answer?
Which of the following are the types of bioreactors?
(i) Simple stirred-tank bioreactor
(ii) Complex Stirred-tank bioreactor
(iii) Sparged stirred-tank bioreactor
(iv) Agitator stirred-tank bioreactor
(i) Simple stirred-tank bioreactor
(ii) Complex Stirred-tank bioreactor
(iii) Sparged stirred-tank bioreactor
(iv) Agitator stirred-tank bioreactor
a)
(i) and (iii)
b)
Only (iii)
c)
(i) and (ii)
d)
(i) and (ii) and (iv)
| | Raghav Bansal answered |
The most commonly used bioreactors are of stirring type. Stirring type bioreactors are simple stirred-tank bioreactor and sparged stirred-tank bioreactor.
Enzyme' Taq polymerase' used in PCR, has been isolated from bacterium ________.- a)Agrobacterium tumefaciens
- b)Thermus aquaticus
- c)Streptomyces albus
- d)Escherichia coli
Correct answer is option 'B'. Can you explain this answer?
Enzyme' Taq polymerase' used in PCR, has been isolated from bacterium ________.
a)
Agrobacterium tumefaciens
b)
Thermus aquaticus
c)
Streptomyces albus
d)
Escherichia coli
| | Dev Patel answered |
The final step of PCR is extension, wherein TaqDNA polymerase (isolated from a thermophilic bacterium Thermus aquaticus) synthesies the DNA region between the primers, using DNTPs (denoxynucleoside triphosphates) and Mg2+. The primers are extened towards each other so that the DNA segment lying between the two primers is copied. The optimum temperature for this polymerisation step is 72∘C. Taq polymerase remains active during high temperature induced denaturation of double stranded DNA.
Primers are- a)Chemically synthesised oligonucleotides that are complementary to the regions of DNA
- b)Chemically synthesised oligonucleotides that are not complementary to the regions of DNA
- c)Chemically synthesised, autonomously replicating circular DNA molecules
- d)Specific sequences present on recombinant DNA
Correct answer is option 'A'. Can you explain this answer?
Primers are
a)
Chemically synthesised oligonucleotides that are complementary to the regions of DNA
b)
Chemically synthesised oligonucleotides that are not complementary to the regions of DNA
c)
Chemically synthesised, autonomously replicating circular DNA molecules
d)
Specific sequences present on recombinant DNA
| | Suresh Iyer answered |
Primers are small, chemically synthesised oligonucleotides that are complementary to the sequences, present at 3' end of the template DNA. They hybridise to the target DNA region, one to each strand of the double helix. These primers are oriented with their ends facing each other allowing synthesis of the DNA towards one another.
Which of the following processes/techniques can be included under biotechnology?
(i) In vitro fertilisation
(ii) Synthesis of a gene
(iii) Correcting a defective gene
(iv) Developing a DNA vaccine- a)(i) and (ii)
- b)(ii) and (iii)
- c)(iii) and (iv)
- d)(i), (ii), (iii) and (iv)
Correct answer is option 'D'. Can you explain this answer?
Which of the following processes/techniques can be included under biotechnology?
(i) In vitro fertilisation
(ii) Synthesis of a gene
(iii) Correcting a defective gene
(iv) Developing a DNA vaccine
(i) In vitro fertilisation
(ii) Synthesis of a gene
(iii) Correcting a defective gene
(iv) Developing a DNA vaccine
a)
(i) and (ii)
b)
(ii) and (iii)
c)
(iii) and (iv)
d)
(i), (ii), (iii) and (iv)
| | Lavanya Menon answered |
Biotechnology deals with techniques of using live micro-organisms, plant or animal cells or their components or enzymes from organisms to produce products and processes (service) useful to human beings. In vitro fertilisation, synthesis of recombinant gene, correcting a defective gene and developing a DNA vaccine are all parts of biotechnology.
The term 'chemical knife' refers to- a)Polymerases
- b)Endonucleases
- c)Ribonudeases
- d)Cellulases
Correct answer is option 'B'. Can you explain this answer?
The term 'chemical knife' refers to
a)
Polymerases
b)
Endonucleases
c)
Ribonudeases
d)
Cellulases
| | Diya Khanna answered |
Explanation:
The term "chemical knife" refers to endonucleases. Endonucleases are a group of enzymes that cleave the phosphodiester bond within a DNA or RNA molecule. They are called "chemical knives" because they can cut DNA or RNA at specific target sequences, similar to how a knife can cut through a specific point on an object.
Endonucleases:
Endonucleases are enzymes that cleave the phosphodiester bond within a DNA or RNA molecule. They are essential for various biological processes like DNA replication, repair, and recombination. Endonucleases can recognize specific nucleotide sequences and cleave the DNA or RNA at those sites.
Types of Endonucleases:
There are several types of endonucleases, including restriction endonucleases, homing endonucleases, and RNA endonucleases. Each type of endonuclease has its own specific function and target sequence.
- Restriction Endonucleases: These enzymes are commonly found in bacteria and are used as a defense mechanism against foreign DNA. They recognize specific sequences of DNA and cleave the DNA at those sites, preventing the foreign DNA from replicating.
- Homing Endonucleases: These enzymes are involved in genetic recombination and can recognize and cleave specific target sequences within DNA. They are often used in genetic engineering techniques to introduce new DNA sequences into an organism.
- RNA Endonucleases: These enzymes cleave RNA molecules at specific sites. They are involved in various processes, including RNA splicing, RNA degradation, and RNA maturation.
Applications of Endonucleases:
Endonucleases have several important applications in molecular biology and biotechnology:
- Genetic Engineering: Endonucleases are used to cleave DNA at specific sites, allowing for the insertion or removal of specific DNA sequences. This is commonly used in genetic engineering techniques like gene cloning and genome editing.
- Diagnostic Tools: Certain endonucleases are used as diagnostic tools to detect specific DNA sequences. For example, restriction endonucleases are often used in DNA fingerprinting techniques.
- Gene Therapy: Endonucleases like CRISPR-Cas9 are being used in gene therapy to correct genetic mutations by editing the DNA sequence.
In conclusion, the term "chemical knife" refers to endonucleases, which are enzymes that cleave DNA or RNA at specific target sequences. They have various applications in molecular biology and biotechnology.
The term "chemical knife" refers to endonucleases. Endonucleases are a group of enzymes that cleave the phosphodiester bond within a DNA or RNA molecule. They are called "chemical knives" because they can cut DNA or RNA at specific target sequences, similar to how a knife can cut through a specific point on an object.
Endonucleases:
Endonucleases are enzymes that cleave the phosphodiester bond within a DNA or RNA molecule. They are essential for various biological processes like DNA replication, repair, and recombination. Endonucleases can recognize specific nucleotide sequences and cleave the DNA or RNA at those sites.
Types of Endonucleases:
There are several types of endonucleases, including restriction endonucleases, homing endonucleases, and RNA endonucleases. Each type of endonuclease has its own specific function and target sequence.
- Restriction Endonucleases: These enzymes are commonly found in bacteria and are used as a defense mechanism against foreign DNA. They recognize specific sequences of DNA and cleave the DNA at those sites, preventing the foreign DNA from replicating.
- Homing Endonucleases: These enzymes are involved in genetic recombination and can recognize and cleave specific target sequences within DNA. They are often used in genetic engineering techniques to introduce new DNA sequences into an organism.
- RNA Endonucleases: These enzymes cleave RNA molecules at specific sites. They are involved in various processes, including RNA splicing, RNA degradation, and RNA maturation.
Applications of Endonucleases:
Endonucleases have several important applications in molecular biology and biotechnology:
- Genetic Engineering: Endonucleases are used to cleave DNA at specific sites, allowing for the insertion or removal of specific DNA sequences. This is commonly used in genetic engineering techniques like gene cloning and genome editing.
- Diagnostic Tools: Certain endonucleases are used as diagnostic tools to detect specific DNA sequences. For example, restriction endonucleases are often used in DNA fingerprinting techniques.
- Gene Therapy: Endonucleases like CRISPR-Cas9 are being used in gene therapy to correct genetic mutations by editing the DNA sequence.
In conclusion, the term "chemical knife" refers to endonucleases, which are enzymes that cleave DNA or RNA at specific target sequences. They have various applications in molecular biology and biotechnology.
The term 'molecular scissors' refers to- a)Recombinant DNA
- b)Restriction enzymes
- c)Taq polymerase
- d)Palindromic nucleotide sequences
Correct answer is option 'B'. Can you explain this answer?
The term 'molecular scissors' refers to
a)
Recombinant DNA
b)
Restriction enzymes
c)
Taq polymerase
d)
Palindromic nucleotide sequences
| | Sounak Shah answered |
The term molecular scissors refers to Restriction enzymes.
Explanation:
Restriction enzymes, also known as restriction endonucleases, are enzymes that can recognize specific DNA sequences and cleave the DNA at those sequences. These enzymes act as "molecular scissors" by cutting the DNA into smaller fragments.
Function of Restriction Enzymes:
Restriction enzymes are naturally occurring enzymes found in bacteria and archaea. They play a crucial role in the defense mechanisms of these organisms by cutting foreign DNA, such as viral DNA, at specific recognition sites. This helps protect the organism from invasion by foreign genetic material.
Recognition Sites:
Restriction enzymes recognize specific DNA sequences called recognition sites or restriction sites. These recognition sites are usually palindromic, meaning they read the same forward and backward on complementary DNA strands. For example, the recognition site for the restriction enzyme EcoRI is 5'-GAATTC-3', which is palindromic.
Cutting the DNA:
Once the restriction enzyme recognizes its specific recognition site, it binds to the DNA at that site and makes a double-stranded cut. The cut can be blunt or staggered, depending on the type of restriction enzyme. Some enzymes create overhangs known as sticky ends, while others create blunt ends.
Applications:
Restriction enzymes are widely used in molecular biology and biotechnology. Their ability to cut DNA at specific sequences allows scientists to manipulate and study DNA in various ways. Some applications of restriction enzymes include:
1. DNA Fragmentation: Restriction enzymes are used to cut DNA into smaller fragments, which can be further analyzed or used in other experiments.
2. DNA Cloning: Restriction enzymes are used to generate compatible ends on DNA fragments and vectors, allowing them to be ligated together during the cloning process.
3. Genetic Engineering: Restriction enzymes are used to insert or remove specific DNA sequences during the process of genetic engineering.
4. DNA Fingerprinting: Restriction enzymes are used in DNA fingerprinting techniques to generate unique DNA fragment patterns for identification purposes.
In summary, the term "molecular scissors" refers to restriction enzymes because they have the ability to recognize specific DNA sequences and cleave the DNA at those sequences. These enzymes play a crucial role in DNA manipulation and various applications in molecular biology and biotechnology.
Explanation:
Restriction enzymes, also known as restriction endonucleases, are enzymes that can recognize specific DNA sequences and cleave the DNA at those sequences. These enzymes act as "molecular scissors" by cutting the DNA into smaller fragments.
Function of Restriction Enzymes:
Restriction enzymes are naturally occurring enzymes found in bacteria and archaea. They play a crucial role in the defense mechanisms of these organisms by cutting foreign DNA, such as viral DNA, at specific recognition sites. This helps protect the organism from invasion by foreign genetic material.
Recognition Sites:
Restriction enzymes recognize specific DNA sequences called recognition sites or restriction sites. These recognition sites are usually palindromic, meaning they read the same forward and backward on complementary DNA strands. For example, the recognition site for the restriction enzyme EcoRI is 5'-GAATTC-3', which is palindromic.
Cutting the DNA:
Once the restriction enzyme recognizes its specific recognition site, it binds to the DNA at that site and makes a double-stranded cut. The cut can be blunt or staggered, depending on the type of restriction enzyme. Some enzymes create overhangs known as sticky ends, while others create blunt ends.
Applications:
Restriction enzymes are widely used in molecular biology and biotechnology. Their ability to cut DNA at specific sequences allows scientists to manipulate and study DNA in various ways. Some applications of restriction enzymes include:
1. DNA Fragmentation: Restriction enzymes are used to cut DNA into smaller fragments, which can be further analyzed or used in other experiments.
2. DNA Cloning: Restriction enzymes are used to generate compatible ends on DNA fragments and vectors, allowing them to be ligated together during the cloning process.
3. Genetic Engineering: Restriction enzymes are used to insert or remove specific DNA sequences during the process of genetic engineering.
4. DNA Fingerprinting: Restriction enzymes are used in DNA fingerprinting techniques to generate unique DNA fragment patterns for identification purposes.
In summary, the term "molecular scissors" refers to restriction enzymes because they have the ability to recognize specific DNA sequences and cleave the DNA at those sequences. These enzymes play a crucial role in DNA manipulation and various applications in molecular biology and biotechnology.
An advantage of using yeasts rather than bacteria as recipient cells for the recombinant DNA of eukaryotes is that yeasts can __________.- a)Produce restriction enzymes
- b)Excise introns from the RNA transcript
- c)Remove methyl groups
- d)Reproduce more rapidly
Correct answer is option 'B'. Can you explain this answer?
An advantage of using yeasts rather than bacteria as recipient cells for the recombinant DNA of eukaryotes is that yeasts can __________.
a)
Produce restriction enzymes
b)
Excise introns from the RNA transcript
c)
Remove methyl groups
d)
Reproduce more rapidly
| | Sinjini Das answered |
Advantages of using yeasts rather than bacteria as recipient cells for the recombinant DNA of eukaryotes:
There are several advantages to using yeasts as recipient cells for the recombinant DNA of eukaryotes, as compared to bacteria. One of the main advantages is that yeasts can excise introns from the RNA transcript.
1. Yeasts can excise introns from the RNA transcript:
- In eukaryotes, genes contain non-coding regions called introns, which need to be removed from the RNA transcript before translation into proteins.
- Bacteria lack the necessary machinery to remove introns, making them unsuitable for processing eukaryotic genes.
- On the other hand, yeasts, being eukaryotic organisms themselves, possess the necessary enzymes and machinery to accurately excise introns from RNA transcripts.
- This ability allows yeasts to properly process and express eukaryotic genes, making them ideal recipient cells for recombinant DNA work involving eukaryotic genes.
2. Other advantages:
- Yeasts are also advantageous because they can carry out post-translational modifications that bacteria cannot perform. These modifications include proper protein folding, glycosylation, phosphorylation, acetylation, and more.
- Yeasts are capable of performing complex cellular processes similar to those found in higher eukaryotes, making them more suitable for expressing and producing eukaryotic proteins.
- Yeasts are more similar to higher eukaryotes in terms of gene regulation and expression, making them a better model system for studying eukaryotic gene function and regulation.
- Yeasts have a longer lifespan and can undergo multiple rounds of cell division, allowing for the production of larger quantities of recombinant proteins compared to bacteria, which have a shorter lifespan and slower reproduction rate.
- Yeasts can also be easily cultured in large-scale fermentation systems, making them suitable for industrial production of recombinant proteins.
In conclusion, yeasts are advantageous over bacteria as recipient cells for recombinant DNA of eukaryotes due to their ability to accurately excise introns from RNA transcripts, perform post-translational modifications, resemble higher eukaryotes in terms of gene regulation, and produce larger quantities of recombinant proteins.
There are several advantages to using yeasts as recipient cells for the recombinant DNA of eukaryotes, as compared to bacteria. One of the main advantages is that yeasts can excise introns from the RNA transcript.
1. Yeasts can excise introns from the RNA transcript:
- In eukaryotes, genes contain non-coding regions called introns, which need to be removed from the RNA transcript before translation into proteins.
- Bacteria lack the necessary machinery to remove introns, making them unsuitable for processing eukaryotic genes.
- On the other hand, yeasts, being eukaryotic organisms themselves, possess the necessary enzymes and machinery to accurately excise introns from RNA transcripts.
- This ability allows yeasts to properly process and express eukaryotic genes, making them ideal recipient cells for recombinant DNA work involving eukaryotic genes.
2. Other advantages:
- Yeasts are also advantageous because they can carry out post-translational modifications that bacteria cannot perform. These modifications include proper protein folding, glycosylation, phosphorylation, acetylation, and more.
- Yeasts are capable of performing complex cellular processes similar to those found in higher eukaryotes, making them more suitable for expressing and producing eukaryotic proteins.
- Yeasts are more similar to higher eukaryotes in terms of gene regulation and expression, making them a better model system for studying eukaryotic gene function and regulation.
- Yeasts have a longer lifespan and can undergo multiple rounds of cell division, allowing for the production of larger quantities of recombinant proteins compared to bacteria, which have a shorter lifespan and slower reproduction rate.
- Yeasts can also be easily cultured in large-scale fermentation systems, making them suitable for industrial production of recombinant proteins.
In conclusion, yeasts are advantageous over bacteria as recipient cells for recombinant DNA of eukaryotes due to their ability to accurately excise introns from RNA transcripts, perform post-translational modifications, resemble higher eukaryotes in terms of gene regulation, and produce larger quantities of recombinant proteins.
During isolation of genetic material, the chemical used to precipitate out the purified DNA is- a)Bromophenol blue
- b)Chilled ethanol
- c)Ethidium bromide
- d)Both A and C
Correct answer is option 'B'. Can you explain this answer?
During isolation of genetic material, the chemical used to precipitate out the purified DNA is
a)
Bromophenol blue
b)
Chilled ethanol
c)
Ethidium bromide
d)
Both A and C
| | Dev Patel answered |
The purified DNA, after treatment with various enzymes, precipitates out after the addition of chilled ethanol.
This is viewed as a collection of fine threads in the suspension and is easily collected. The process is known as DNA spooling.
This is viewed as a collection of fine threads in the suspension and is easily collected. The process is known as DNA spooling.
In addition to the Taq polymerase enzyme, which other thermostable DNA polymerases have been isolated to be used in a polymerase chain reaction (PCR)?- a)Pfu polymerase isolated from Pyrococcus furiosus
- b)Tli polymerase (vent polymerase) isolated from Thermococcus litoralis
- c)Both (a) and (b)
- d)None of these
Correct answer is option 'C'. Can you explain this answer?
In addition to the Taq polymerase enzyme, which other thermostable DNA polymerases have been isolated to be used in a polymerase chain reaction (PCR)?
a)
Pfu polymerase isolated from Pyrococcus furiosus
b)
Tli polymerase (vent polymerase) isolated from Thermococcus litoralis
c)
Both (a) and (b)
d)
None of these
| | Hansa Sharma answered |
In addition to Tag DNA polymerase, Pfu polymerase and Tli polymerase have been isolated which are also thermostable. Pfu polymerase is isolated from Pyrococcus furiosus. Tli (vent) polymerase is isolated from Thermococcus litoralis.
Genetic engineering is possible, because- a)We can cut DNA at specific sites by endonucleases
- b)Restriction endonucleases purified from bacteria can be used in vitro
- c)The phenomenon of transduction in bacteria is well understood
- d)We can see DNA by electron microscope
Correct answer is option 'A'. Can you explain this answer?
Genetic engineering is possible, because
a)
We can cut DNA at specific sites by endonucleases
b)
Restriction endonucleases purified from bacteria can be used in vitro
c)
The phenomenon of transduction in bacteria is well understood
d)
We can see DNA by electron microscope
| | Riya Banerjee answered |
Genetic engineering is the artificail synthesis, isolation modification, cimbination, addition and repaor of the genetic material (DNA) to alter the phenotype of the host organism to suit human needs. It is the manipulation of genes by man in vitro. Restriction endonucleases play major role in gentic engineering as they can cut DNA at specific sites.
Stirred-tank bioreactors have advantages over shake flasks because they ___________.- a)Provide high temperature and pH
- b)Provide better aeration and mixing properties
- c)Do not allow the entry of CO2
- d)Are easy to operate
Correct answer is option 'B'. Can you explain this answer?
Stirred-tank bioreactors have advantages over shake flasks because they ___________.
a)
Provide high temperature and pH
b)
Provide better aeration and mixing properties
c)
Do not allow the entry of CO2
d)
Are easy to operate
| | Dev Patel answered |
Stirred-tank bioreactor is used for processing large volumes of culture. It is a cylindrical tank with a curved base to facilitate the mixing of the reactor contents. The stirrer facilitates even mixing and oxygen availability throughout the bioreactor.
Eukaryotic genes do not function properly when cloned into a bacterial cell because- a)of the inability to excise introns and destruction by bacterial restriction enzymes
- b)of high pH present in bacterial cells
- c)of inappropriate insertion of genes
- d)both (a) and (b)
Correct answer is option 'A'. Can you explain this answer?
Eukaryotic genes do not function properly when cloned into a bacterial cell because
a)
of the inability to excise introns and destruction by bacterial restriction enzymes
b)
of high pH present in bacterial cells
c)
of inappropriate insertion of genes
d)
both (a) and (b)
| | Mira Joshi answered |
Eukaryotic genes do not function properly When transferred into bacterial cell because introns are present eukaryotic cells but are absent in prokaryotic cells. Hence, when bacterial cell is transformed with recombinant DNA is genrated human gene, it could not process it. As a result no desired protien will be produced.
Who is the father of genetic engineering?- a)Steward Linn
- b)Stanley Cohen
- c)Paul Berg
- d)Kary Mullis
Correct answer is option 'C'. Can you explain this answer?
Who is the father of genetic engineering?
a)
Steward Linn
b)
Stanley Cohen
c)
Paul Berg
d)
Kary Mullis
| | Dev Patel answered |
In 1972, genetic engineering was started by Paul Berg. He was able to introduce a gene of the SV-40 virus into a bacterium with the help of lambda phage. Berg is often considered as "Father of genetic engineering". He was awarded Nobel Prize in 1980.
The different steps of recombinant DNA technology are given below randomly.
(i) Isolation of the DNA fragments or genes to be cloned
(ii) Introduction of the recombinant DNA into a suitable cell (usually E.coli) called host (transformation)
(iii) Multiplication/expression of the introduced gene in the host
(iv) Selection of the transformed host cells, and identification of the clone containing the desired gene/DNA fragment
(v) Insertion of the isolated gene in a suitable plasmid vectorWhich of the following represents the correct sequence of steps?- a)(i), (iii), (ii), (iv), (v)
- b)(iii), (ii), (i), (v), (iv)
- c)(i), (v), (ii), (iv), (iii)
- d)(v), (i), (iii), (iv), (ii)
Correct answer is option 'C'. Can you explain this answer?
The different steps of recombinant DNA technology are given below randomly.
(i) Isolation of the DNA fragments or genes to be cloned
(ii) Introduction of the recombinant DNA into a suitable cell (usually E.coli) called host (transformation)
(iii) Multiplication/expression of the introduced gene in the host
(iv) Selection of the transformed host cells, and identification of the clone containing the desired gene/DNA fragment
(v) Insertion of the isolated gene in a suitable plasmid vector
(i) Isolation of the DNA fragments or genes to be cloned
(ii) Introduction of the recombinant DNA into a suitable cell (usually E.coli) called host (transformation)
(iii) Multiplication/expression of the introduced gene in the host
(iv) Selection of the transformed host cells, and identification of the clone containing the desired gene/DNA fragment
(v) Insertion of the isolated gene in a suitable plasmid vector
Which of the following represents the correct sequence of steps?
a)
(i), (iii), (ii), (iv), (v)
b)
(iii), (ii), (i), (v), (iv)
c)
(i), (v), (ii), (iv), (iii)
d)
(v), (i), (iii), (iv), (ii)
| | Pallavi Pillai answered |
Understanding Recombinant DNA Technology Steps
Recombinant DNA technology involves several crucial steps to successfully clone and express a gene of interest. The correct sequence of these steps is pivotal for achieving the desired outcome.
1. Isolation of the DNA Fragments
- This is the first step where the specific DNA fragments or genes that need to be cloned are isolated from the source organism.
2. Insertion into a Suitable Plasmid Vector
- The isolated gene is then inserted into a suitable plasmid vector. This vector is essential as it carries the gene into the host cell.
3. Introduction of Recombinant DNA into Host Cells
- The next step involves the introduction of the recombinant DNA (plasmid containing the gene) into a host cell, typically E. coli. This process is known as transformation.
4. Selection of Transformed Host Cells
- After transformation, it is crucial to select the host cells that have successfully taken up the recombinant DNA. Identification techniques are used to isolate those clones containing the desired gene.
5. Multiplication/Expression of the Introduced Gene
- Finally, the transformed cells multiply and express the introduced gene, producing the desired protein or product.
Correct Sequence Summary
- The correct sequence of steps is:
- (i) Isolation of the DNA fragments
- (v) Insertion of the isolated gene in a suitable plasmid vector
- (ii) Introduction of the recombinant DNA into a suitable cell (transformation)
- (iv) Selection of the transformed host cells
- (iii) Multiplication/expression of the introduced gene
This systematic approach ensures that the recombinant DNA technology is executed effectively, leading to successful gene cloning and expression.
Recombinant DNA technology involves several crucial steps to successfully clone and express a gene of interest. The correct sequence of these steps is pivotal for achieving the desired outcome.
1. Isolation of the DNA Fragments
- This is the first step where the specific DNA fragments or genes that need to be cloned are isolated from the source organism.
2. Insertion into a Suitable Plasmid Vector
- The isolated gene is then inserted into a suitable plasmid vector. This vector is essential as it carries the gene into the host cell.
3. Introduction of Recombinant DNA into Host Cells
- The next step involves the introduction of the recombinant DNA (plasmid containing the gene) into a host cell, typically E. coli. This process is known as transformation.
4. Selection of Transformed Host Cells
- After transformation, it is crucial to select the host cells that have successfully taken up the recombinant DNA. Identification techniques are used to isolate those clones containing the desired gene.
5. Multiplication/Expression of the Introduced Gene
- Finally, the transformed cells multiply and express the introduced gene, producing the desired protein or product.
Correct Sequence Summary
- The correct sequence of steps is:
- (i) Isolation of the DNA fragments
- (v) Insertion of the isolated gene in a suitable plasmid vector
- (ii) Introduction of the recombinant DNA into a suitable cell (transformation)
- (iv) Selection of the transformed host cells
- (iii) Multiplication/expression of the introduced gene
This systematic approach ensures that the recombinant DNA technology is executed effectively, leading to successful gene cloning and expression.
Match the scientists in column I with their related discoveries in column II and select the correct option from the given codes.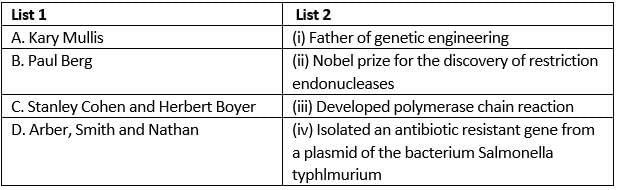
- a)A−(iii), B−(i), C−(iv), D−(ii)
- b)A−(iii), B−(iv), C−(i), D−(ii)
- c)A−(iv), B−(ii), C−(iii), D−(i)
- d)A−(i), B−(iii), C−(iv), D−(ii)
Correct answer is option 'A'. Can you explain this answer?
Match the scientists in column I with their related discoveries in column II and select the correct option from the given codes.

a)
A−(iii), B−(i), C−(iv), D−(ii)
b)
A−(iii), B−(iv), C−(i), D−(ii)
c)
A−(iv), B−(ii), C−(iii), D−(i)
d)
A−(i), B−(iii), C−(iv), D−(ii)
| | Priya Menon answered |
- The father of genetic engineering is Paul Berg. He creates the recombinant DNA technology in 1972.
- Kary Mullis invents the polymerase chain reaction in 1993. Arber, Smith and Nathan discovered the restriction endonuclease enzymes in 1978.
- Stanley Cohen and Herbert Boyer constructed artificial recombinant DNA for the first time. They got the idea of linking a gene encoding for antibiotic resistance with a native plasmid of Salmonella typhinurium.
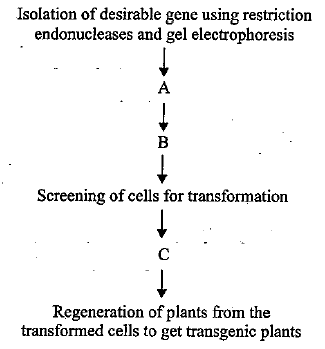
The given flow chart depicts the steps to transfer a desirable gene of interest into a plant.
Identify the missing steps (A, B and C) with regard to following statements and select the correct option.
(i) Joining of desirable gene to a suitable cloning vector using ligases to create a recombinant DNA molecule
(ii) Selection of transformed cells
(iii) Transferring the recombinant DNA molecules to the target cells
- a)A - i, B - ii, C - iii
- b)A - i, B - iii, C - ii
- c)A - ii, B - iii, C - i
- d)A - iii, B - i, C - ii
Correct answer is option 'B'. Can you explain this answer?
The given flow chart depicts the steps to transfer a desirable gene of interest into a plant.
Identify the missing steps (A, B and C) with regard to following statements and select the correct option.
(i) Joining of desirable gene to a suitable cloning vector using ligases to create a recombinant DNA molecule
(ii) Selection of transformed cells
(iii) Transferring the recombinant DNA molecules to the target cells

Identify the missing steps (A, B and C) with regard to following statements and select the correct option.
(i) Joining of desirable gene to a suitable cloning vector using ligases to create a recombinant DNA molecule
(ii) Selection of transformed cells
(iii) Transferring the recombinant DNA molecules to the target cells
a)
A - i, B - ii, C - iii
b)
A - i, B - iii, C - ii
c)
A - ii, B - iii, C - i
d)
A - iii, B - i, C - ii
| | Raghav Bansal answered |
Steps to transfer a desirable gene of interest into a plant:
- Isolation of desirable gene using restriction endonucleases and gel electrophoresis.
- Joining of desirable gene to a suitable cloning vector using ligases to create a recombinant DNA molecule.
- Transferring the recombinant DNA molecules to the target cells.
- Screening of cells.
- Transformation.
- Selection of transformed cells.
- Regeneration of plants from the transformed cells to get transgenic plants.
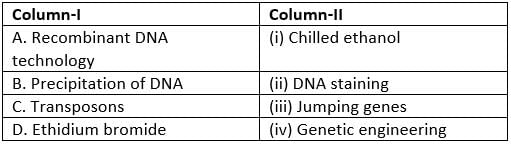
Match column I with column II and select the correct answer from the given codes.
- a)A-(iv), B-(i), C-(iii), D-(ii)
- b)A-(i), B-(iii), C-(ii), D-(iv)
- c)A-(ii), B-(i), C-(iii), D-(iv)
- d)A-(iv), B-(ii), C-(i), D-(iii)
Correct answer is option 'A'. Can you explain this answer?
Match column I with column II and select the correct answer from the given codes.


a)
A-(iv), B-(i), C-(iii), D-(ii)
b)
A-(i), B-(iii), C-(ii), D-(iv)
c)
A-(ii), B-(i), C-(iii), D-(iv)
d)
A-(iv), B-(ii), C-(i), D-(iii)
 | Dr. Mohit Rajpoot answered |
Correct answer: A
A → (iv): Recombinant DNA technology involves joining DNA from different sources to create new combinations and is commonly termed genetic engineering.
B → (i): DNA is commonly recovered from solution by precipitation with chilled ethanol; the cold ethanol causes DNA to aggregate and precipitate out.
C → (iii): Transposons are mobile genetic elements known as jumping genes because they can move from one chromosomal location to another.
D → (ii): Ethidium bromide is an intercalating chemical used for DNA staining in gel electrophoresis; it binds DNA and fluoresces under UV light.
A-(iv), B-(i), C-(iii), D-(ii) is therefore the correct matching, which corresponds to option A.
Chapter doubts & questions for Biotechnology and Its Processes - 30-Day Revision Course for NEET 2026 is part of NEET exam preparation. The chapters have been prepared according to the NEET exam syllabus. The Chapter doubts & questions, notes, tests & MCQs are made for NEET 2026 Exam. Find important definitions, questions, notes, meanings, examples, exercises, MCQs and online tests here.
Chapter doubts & questions of Biotechnology and Its Processes - 30-Day Revision Course for NEET in English & Hindi are available as part of NEET exam. Download more important topics, notes, lectures and mock test series for NEET Exam by signing up for free.
;
Signup on EduRev and stay on top of your study goals
10M+ students crushing their study goals daily

