Proteins - Biomolecules | Organic Chemistry PDF Download
(A) Classification and Chemistry of Protein Structure
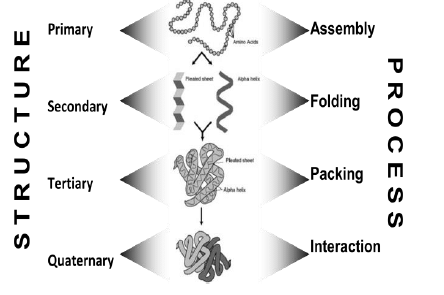
a) Protein Assembly
Protein assembly occurs at ribosomes. It involves dehydration synthesis and polymerization of amino acids attached to t-RNA and yields primary structure.
Primary Structure
It is linear, ordered and one dimensional. It is by convention, written from amino end to carboxyl end. A perfectly linear amino acid polymer is neither functional nor energetically favorable → folding!
b.) Protein Folding
It occurs in the cytosol and involves localized spatial interaction among primary structure elements, i.e. the amino acids. It may or may not involve chaperone proteins. It tumbles towards conformations that reduce ΔE (this process is thermo-dynamically favorable) and yields secondary structure.
Secondary Structure
Secondary structure is non-linear and 3 dimensional. It localized to regions of an amino acid chain. It is formed and stabilized by hydrogen bonding, electrostatic and van der Waals interactions.
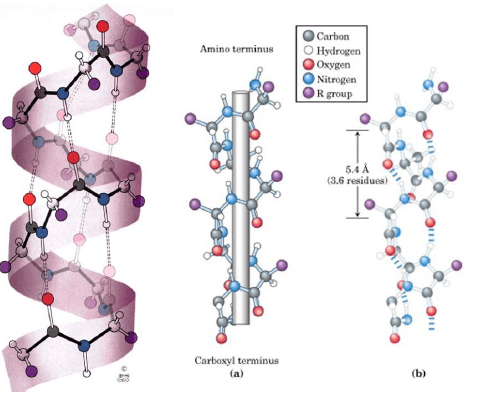
c.) Protein Packing
Protein packing occurs in the cytosol (~60% bulk water, ~40% water of hydration). It involves interaction between secondary structure elements and solvent. It may be promoted by chaperones, membrane proteins and tumbles into molten globule states. Overall entropy loss is small enough so enthalpy determines sign of ΔE, which decreases (loss in entropy from packing counteracted by gain from desolvation and reorganization of water, i.e. hydrophobic effect). Hence, yields tertiary structure.
Tertiary Structure
Tertiary structure is non-linear and 3-dimensional. It is global but restricted to the amino acid polymer. It is formed and stabilized by hydrogen bonding, covalent (e.g. disulfide) bonding, hydrophobic packing toward core and hydrophilic exposure to solvent.
A globular amino acid polymer folded and compacted is somewhat functional (catalytic) and energetically favorable → interaction!
d.) Protein Interaction
Protein interaction occurs in the cytosol, in close proximity to other folded and packed proteins. It involves interaction among tertiary structure elements of separate polymer chains. It may be promoted by chaperones, membrane proteins, cytosolic and extracellular elements as well as the proteins’ own propensities. ΔE decreases further due to further desolvation and reduction of surface area
1. Globular proteins, e.g. hemoglobin, largely involved in catalytic roles
2. Fibrous proteins, e.g. collagen, largely involved in structural roles
Hence, yields quaternary structure.
Quaternary Structure
Quaternary structure is non-linear and 3 dimensional. It is global, and across distinct amino acid polymers. It is formed by hydrogen bonding, covalent bonding, hydrophobic packing and hydrophilic exposure. It is favorable, functional structures occur frequently and have been categorized.
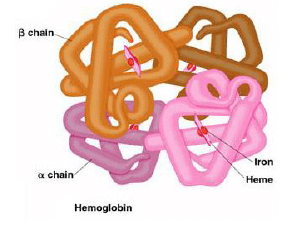
(B) Classification based on protein function
1. Catalytic: Rubisco
2. Communication: Hormones (eg insulin) and Neurotransmitters
3. Protective: Immunoglobulin, Fibrinogen, Blood clotting factors
4. Structural: Collagen, Spiders silk
5. Pigments: Rhodopsin
6. Storage: Ovalbumen (in eggs), Casein (in milk)
7. Transport: Hemoglobin
8. Contractile: Actin, Myosin
9. Toxins: Snake venom
Denaturation of proteins
The three-dimensional conformation of proteins is stabilized by bonds or interactions between R groups of amino acids within the molecule. Most of these bonds and interactions are relatively weak and they can be disrupted or broken. This results in a change to the conformation of the protein, which is called denaturation.
A denatured protein does not normally return to its former structure – the denaturation is permanent. Soluble proteins often become insoluble and form a precipitate.
Heat can cause denaturation: vibrations within the molecule breaks intramolecular bonds or interactions.
Extremes of pH can cause denaturation: charges on R groups are changed, breaking ionic bonds within the protein or causing new ionic bonds to form.
Thermophiles are organisms (often archea or eubacteria) that live in relatively hot conditions (45 to122°C). In order that they can survive, their proteins are stable at the higher than normal temperatures they experience.
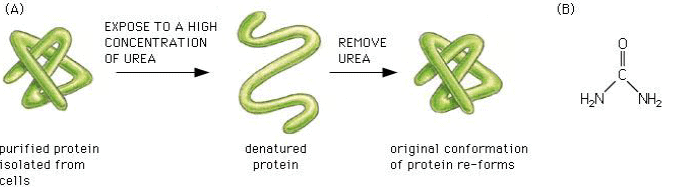
Molecular chaperones are small proteins that help guide the folding and can help keep the new protein from associating with the wrong partner.
Domains
- A domain is a basic structural unit of a protein structure – distinct from those that make up the conformations
- Part of protein that can fold into a stable structure independently
- Different domains can impart different functions to proteins
- Proteins can have one to many domains depending on protein size
Useful Proteins
There are thousands and thousands of different combinations of amino acids that can make up proteins and that would increase if each one had multiple shapes. Proteins usually have only one useful conformation because otherwise it would not be efficient use of the energy available to the system. Natural selection has eliminated proteins that do not perform a specific function in the cell.
Proteins at Work
The conformation of a protein gives it a unique function. To work proteins must interact with other
molecules, usually one or a few molecules from the thousands to one protein.
- Ligand – The molecule that a protein can bind
- Binding site – Part of the protein that interacts with ligand. It consists of a cavity formed by a specific arrangement of amino acids.
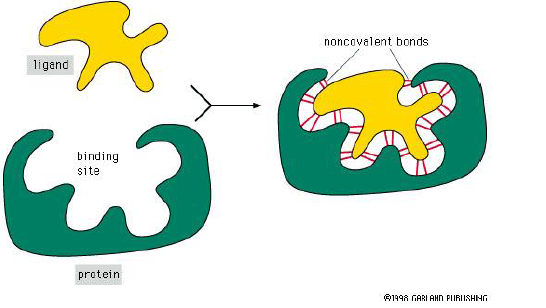
Formation of Binding Site
- The binding site forms when amino acids from within the protein come together in the folding
- The remaining sequences may play a role in regulating the protein’s activity.
Antibody Family
A family of proteins that can be made to bind to almost any molecule. Antibodies (immunoglobulins) are made in response to a foreign molecule ie. bacteria, virus, pollen… called the antigen. They bind together tightly and therefore, inactivates the antigen or marks it for destruction.
Antibodies:
- Y-shaped molecules with 2 binding sites at the upper ends of the Y
- The loops of polypeptides on the end of the binding site are what imparts the recognition of the antigen
- Changes in the sequence of the loops make the antibody recognize different antigens – specificity.
Binding Strength
- Can be measured directly
- Antibodies and antigens are mixing around in a solution, eventually they will bump into each other in a way that the antigen sticks to the antibody, eventually they will separate due to the motion in the molecules
- This process continues until the equilibrium is reached – number sticking is constant and number leaving is constant
- This can be determined for any protein and its ligand.
Equilibrium Constant
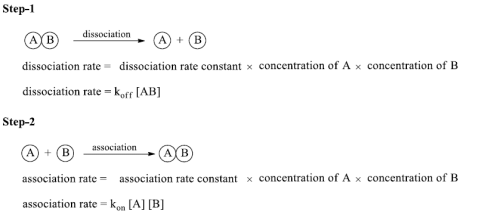
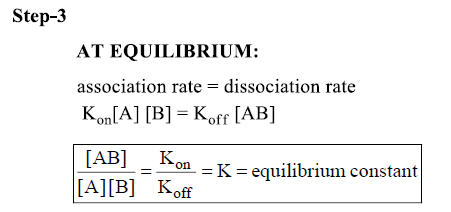


Concentration of antigen, antibody and antigen/antibody complex at equilibrium can be measured-equilibrium constant (K)
- Larger the K the tighter the binding or the more non-covalent bonds that hold the 2 together.
Enzymes as Catalysts
Enzymes are proteins that bind to their ligand as the 1st step in a process. An enzyme’s ligand is called a substrate. It may be one or more molecules. Output of the reaction is called the product. Enzymes can repeat these steps many times and rapidly, called catalysts.
Enzymes at Work
- Lysozyme is an important enzyme that protects us from bacteria by making holes in the bacterial cell wall and causing it to break
- Lysozyme adds H2O to the glycosidic bond in the cell wall
- Lysozyme holds the polysaccharide in a position that allows the H2O to break the bond – this is the transition state – state between substrate and product
- Active site is a special binding site in enzymes where the chemical reaction takes place.
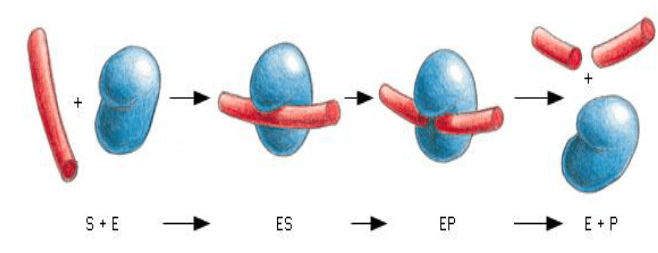
Non-covalent bonds hold the polysaccharide in the active site until the reaction occurs.
Features of Enzyme Catalysis
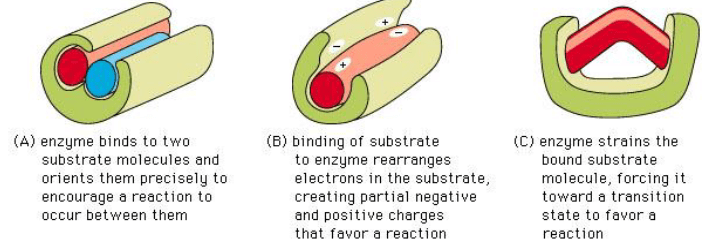
Enzyme Performance
E + S ↔ES ↔ EP ↔ E + P
Step 1: Binding of the substrate
- Limiting step depending on [S] and/or [E]
- Vmax – maximum rate of the reaction
- Turnover number determines how fast the substrate can be processed = rate of rxn ÷ [E]
Step 2: Stabilization of transition state
- State of substrate prior to becoming product
- Enzymes lowers the energy of transition state and therefore accelerates the reaction
Reaction Rates
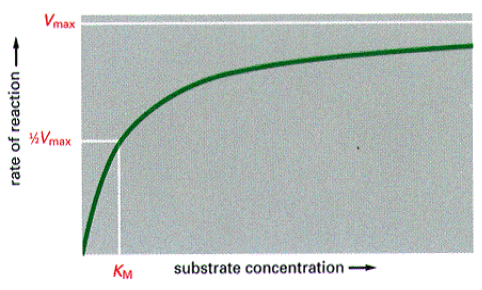
KM – [S] that allows reaction to proceed at 1/2 it maximum rate.
Prosthetic Groups
- Occasionally the sequence of the protein is not enough for the function of the protein
- Some proteins require a non-protein molecule to enhance the performance of the protein
- Hemoglobin requires heme (iron containing compound) to carry the O2
- When a prosthetic group is required by an enzyme it is called a co-enzyme
- Usually a metal or vitamin
- These groups may be covalently or non-covalently linked to the protein.
|
44 videos|102 docs|52 tests
|
FAQs on Proteins - Biomolecules - Organic Chemistry
| 1. What are proteins and why are they considered biomolecules? |  |
| 2. How are proteins formed in living organisms? |  |
| 3. What are the primary functions of proteins in the human body? |  |
| 4. Can proteins be obtained solely from animal sources? |  |
| 5. Are all proteins equally beneficial for the body? |  |





















