NCERT Exemplar: Biotechnology: Principle & Processes | Biology Class 12 - NEET PDF Download
MULTIPLE CHOICE QUESTIONS
Q.1. Rising of dough is due to:
(a) Multiplication of yeast
(b) Production of CO2
(c) Emulsification
(d) Hydrolysis of wheat flour starch into sugars.
Ans. (b)
Solution.
Dough, which is used for making dosa and idli is fermented by bacteria. The puffed up appearance of dough is due to the production of CO2 gas. Dough which is used for making bread, is fermented by fungi Saccharomyces cerevisiae (Baker’s yeast).
Q.2. Which of the following enzymes catalyse the removal of nucleotides from the ends of DNA?
(a) Endonuclease
(b) Exonuclease
(c) DNA ligase
(d) Hind - II
Ans. (b)
Solution.
Exonucleases remove nucleotides from the ends of the DNA whereas endonucleases make cuts at specific positions within the DNA.
Q.3. The transfer of genetic material from one bacterium to another through the mediation of a viral vector is termed as:
(a) Transduction
(b) Conjugation
(c) Transformation
(d) Translation
Ans. (a)
Solution.
The transfer of genetic material from one bacterium to another through the mediation of a vector like virus is termed as transduction.
Q.4. Which of the given statements is correct in the context of visualizing DNA molecules separated by agarose gel electrophoresis?
(a) DNA can be seen in visible light
(b) DNA can be seen without staining in visible light
(c) Ethidium bromide stained DNA can be seen in visible light
(d) Ethidium bromide stained DNA can be seen under exposure to UV light
Ans. (d)
Solution.
The separated DNA fragments can he visualised only after staining the DNA with a compound known as ethidium bromide followed by exposure to ultraviolet radiation (we cannot see pure DNA fragments in the visible light and without staining).
Q.5. 'Restriction' in Restriction enzyme refers to:
(a) Cleaving of phosphodiester bond in DNA by the enzyme
(b) Cutting of DNA at specific position only
(c) Prevention of the multiplication of bacteriophage by the host bacteria
(d) All of the above
Ans. (c)
Solution.
'Restriction’inRestrictionenzymereferstopreventionofthemultiplication of bacteriophage in bacteria.
Q.6. Which of the following is not required in the preparation of a recombinant DNA molecule?
(a) Restriction endonuclease
(b) DNA ligase
(c) DNA fragments
(d) E.coli
Ans. (d)
Solution.
Genetic engineering or r-DNA technology can be accomplished only if we have the key tools:
(1) Restriction enzymes
(2) Polymerase enzymes
(3) Ligases
(4) Vectors
(5) Host organism
Q.7. In agarose gel electrophoresis, DNA molecules are separated on the basis of their:
(a) Charge only
(b) Size only
(c) Charge to size ratio
(d) All of the above
Ans. (d)
Solution.
The differential mobility of DNA depends upon charge and size of DNA.
Q.8. The most important feature in a plasmid to serve as a vector in gene cloning experiment is:
(a) Origin of replication (ori)
(b) Presence of a selectable marker
(c) Presence of sites for restriction endonuclease
(d) Its size
Ans. (a)
Solution.
The most important feature in a plasmid to be used as a vector is origin of replication (ori).
Q.9. While isolating DNA from bacteria, which of the following enzymes is not required?
(a) Lysozyme
(b) Ribonuclease
(c) Deoxyribonuclease
(d) Protease
Ans. (c)
Solution.
While isolating DNA from bacteria lysozyme, ribonuclease and protease, enzymes is used.
Q.10. Which of the following contributed in popularising the PCR (polymerase chain reactions) technique?
(a) Easy availability of DNA template
(b) Availability of synthetic primers
(c) Availability of cheap deoxyribonucleotides
(d) Availability of 'Thermostable' DNA polymerase
Ans. (d)
Solution.
If the process of replication ‘of DNA is repeated many times, the segment of DNA can be amplified to approximately billion times i.e., 1 billion copies are made. Such repeated amplification is achieved by the use of a thermostable DNA polymerase (isolated from a thermophilic bacterium Thermits aquaticus), which is active during the high temperature induced denaturation of double stranded DNA.
Q.11. An antibiotic resistance gene in a vector usually helps in the selection of:
(a) Competent bacterial cells
(b) Transformed bacterial cells
(c) Recombinant bacterial cells
(d) None of the above
Ans. (b)
Solution.
An antibiotic resistance gene in a vector usually helps in the selection of transformed cells.
Q.12. Significance of 'heat shock' method in bacterial transformation is to facilitate:
(a) Binding of DNA to the cell wall
(b) Uptake of DNA through membrane transport proteins
(c) Uptake of DNA through transient pores in the bacterial cell wall
(d) Expression of antibiotic resistance gene
Ans. (c)
Solution.
Significance of ‘heat shock’ method in bacterial transformation is to facilitate uptake of DNA through transient pores in the bacterial cell wall.
Q.13. The role of DNA ligase in the construction of a recombinant DNA molecule is:
(a) Formation of phosphodiester bond between two DNA fragments
(b) Formation of hydrogen bonds between sticky ends of DNA fragments
(c) Ligation of all purine and pyrimidine bases
(d) None of the above
Ans. (a)
Solution.
The role of DNA ligase in the construction of a recombinant DNA molecule is formation of phosphodiester bond between two DNA fragments.
Q.14. Which of the following bacteria is not a source of restriction endonuclease?
(a) Haemophilus influenzae
(b) Escherichia coli
(c) Entamoeba coli
(d) Bacillus amyloliquefaciens
Ans. (c)
Solution.
Haemophilus influenzae, Escherichia coli and Bacillius quifacieus are source of restriction endonuclease.
Q.15. Which of the following steps are catalysed by Taq DNA polymerase in a PCR reaction?
(a) Denaturation of template DNA
(b) Annealing of primers to template DNA
(c) Extension of primer end on the template DNA
(d) All of the above
Ans. (c)
Solution.
Taq polymerase is used between annealing and extension, help in extension of primer end on the template DNA.
Q.16. A bacterial cell was transformed with a recombinant DNA molecule that was generated using a human gene. However, the transformed cells did not produce the desired protein. Reasons could be:
(a) Human gene may have intron which bacteria cannot process
(b) Amino acid codons for humans and bacteria are different
(c) Human protein is formed but degraded by bacteria
(d) All of the above
Ans. (a)
Solution.
A bacterial cell was transformed with a recombinant DNA that was generated using a human gene However, the transformed cells did not produce the desired protein because human gene may have intron which bacteria cannot process.
Q.17. Which of the following should be chosen for best yield if one were to produce a recombinant protein in large amounts?
(a) Laboratory flask of largest capacity
(b) A stirred-tank bioreactor without in-lets and out-lets
(c) A continuous culture system
(d) Any of the above
Ans. (c)
Solution.
- After having cloned the gene of interest and having optimised the conditions to induce the expression of the target protein one has to consider producing it on a large scale. Small volume cultures cannot yield appreciable quantities of products.
- To produce in large quantities the development of bioreactors, where large volume (100-1000 litres) of culture (continuous) can be processed, was required.
Q.18. Who among the following was awarded the Nobel Prize for the development of PCR technique?
(a) Herbert Boyer
(b) Hargovind Khurana
(c) Kary Mullis
(d) Arthur Kornberg
Ans. (c)
Solution.
Kary Mullis was awarded the Nobel Prize for the development of PCR technique.
Q.19. Which of the following statements does not hold true for restriction enzyme?
(a) It recognises a palindromic nucleotide sequence
(b) It is an endonuclease
(c) It is isolated from viruses
(d) It can produce the same kind of sticky ends in different DNA molecules
Ans. (c)
Solution.
Restriction enzyme:
- recognises a palindromic nucleotide sequence
- is an endonuclease isolated from bacteria
- produces the same kind of sticky ends in different DNA molecules
VERY SHORT ANSWER TYPE QUESTIONS
Q.1. How is copy number of the plasmid vector related to yield of recombinant protein?
Ans. Higher the copy number of vector plasmid, higher the copy number of gene and consequently, protein coded by the gene is produced in high amount.
Q.2. Would you choose an exonuclease while producing a recombinant DNA molecule?
Ans. No, as exonuclease acts on the free ends of linear DNA molecule. Therefore, instead of producing DNA fragments with sticky ends, it will shorten or completely degrade the DNA fragment containing the gene of interest, and the circular plasmid (vector) will not get cut as it lacks free ends.
Q.3. What does H in’ ‘d’ and ‘III’ refer to in the enzyme Hind III?
Ans. ‘H’ from name of genus Haemophilus, ‘d’ from name of strain Rd. TIT indicates the order in which the enzyme were isolated from Rd strain of bacteria.
Q.4. Restriction enzymes should not have more than one site of action in the cloning site of a vector. Comment.
Ans. If the restriction enzymes have more than one recognition site in a vector, than the vector itself will get fragmented on treatment with the restriction enzyme.
Q.5. What does ‘competent’ refer to in competent cells used in transformation experiments?
Ans. Competent means bacterial cells, on treatment with CaCl2, are made capable of taking up foreign DNA.
Q.6. What is the significance of adding proteases at the time of isolation of genetic material (DNA).
Ans. Role of proteases is to degrade the proteins present inside a cell (from which DNA is being isolated). If the proteins are not removed from DNA preparation then they could interfere with any downstream treatment of DNA (such action of restriction endonuclease, DNA ligase etc).
Q.7. While doing a PCR, ‘denaturation’ step is missed. What will be its effect on the process?
Ans. If denaturation of double-stranded DNA does not take place, then primers will not be able to anneal to the template, no extension will take place, hence no amplification will occur.
Q.8. Name a recombinant vaccine that is currently being used in vaccination program.
Ans. Hepatitis B recombinant vaccine-engerix is used for vaccination of hepatitis virus.
Q.9. Do biomolecules (DNA, protein) exhibit biological activity in anhydrous conditions?
Ans. No, biomolecules like DNA and protein cannot exhibit biological activity in anhydrous conditions. Hence, life is unsustainable without water.
Q.10. What modification is done on the Ti plasmid of Agrobacterium tumefaciens to convert it into a cloning vector?
Ans. The tumor inducing (Ti) plasmid of Agrobacterium tumefaciens has now been modified (disarmed) into a cloning vector which is no more pathogenic to the plants but is still able to use the mechanisms to deliver genes of our interest into a variety of plants.
SHORT ANSWER TYPE QUESTIONS
Q.1. What is meant by gene cloning?
Ans. Gene cloning refers to a process in which a gene of interest is ligated to a vector. The recombinant DNA thus produced is introduced in a host cell by transformation. Each cell gets one DNA molecule and when the transformed cell grows to a bacterial colony, each cell in the colony has a copy of the gene. This is precisely gene cloning.
Q.2. Both a wine maker and a molecular biologist who had developed a recombinant vaccine claim to be biotechnologists. Who in your opinion is correct?
Ans. Both. As biotechnology is a very wide area which deals with techniques of using a ‘natural’ organism (or its parts) as well as genetically modified organism to produce products and processes useful for mankind. A wine maker employs a strain-of yeast to produce wine by fermentation (a natural phenomenon), while the molecular biologist has cloned gene for the antigen (that is used as vaccine) in an organism which allows the production of the antigen in large amount.
Q.3. A recombinant DNA molecule was created by ligating a gene to a plasmid vector. By mistake, an exonuclease was added to the tube containing the recombinant DNA. How does this affect the next step in the experiment i.e. bacterial transformation?
Ans. The experiment will not likely to be affected as recombinant DNA molecule is circular closed, with no free ends. Hence, it will not be a substrate for exonuclease enzyme which removes nucleotides from the free ends of DNA.
Q.4. Restriction enzymes that are used in the construction of recombinant DNA are endonucleases which cut the DNA at ‘specific-recognition sequence’. What would be the disadvantage if they do not cut the DNA at specific-recognition sequence?
Ans. If the restriction enzymes would cut DNA at random sites instead of at specific sites, then the DNA fragments obtained will not have ‘sticky ends’. In the absence of sticky ends, construction of recombinant DNA molecule would not be possible.
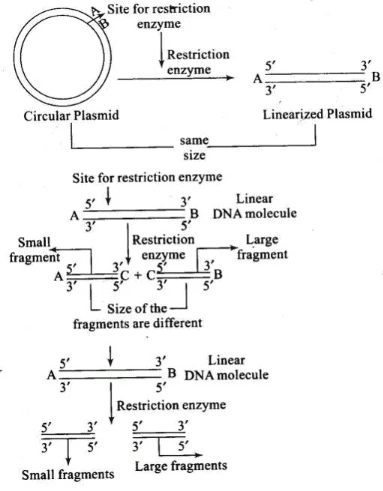
Q.5. A plasmid DNA and a linear DNA (both are of the same size) have one site for a restriction endonuclease. When cut and separated on agarose gel electrophoresis, plasmid shows one DNA band while linear DNA shows two fragments. Explain.
Ans. It is because plasmid is a circular DNA molecule. When cut with enzyme, it becomes linear but does not get fragmented. Whereas, a linear DNA molecule gets cut into two fragments. Hence, a single DNA band is observed for plasmid while two DNA bands are observed for linear DNA in agarose gel.
Q.6. How does one visualise DNA on an agarose gel?
Ans. A compound called Ethidium Bromide stains DNA, which on irradiating with Ultraviolet, fluoresce gives orange light. Hence, DNA fragments appear as orange band in the presence of Ethidium Bromide and UV.
Q.7. A plasmid without a selectable marker was chosen as vector for cloning a gene. How does this affect the experiment?
Ans. In a gene, cloning experiment, first a recombinant DNA molecule is constructed, where the gene of interest is ligated to the vector (the step would not be affected) and introduced inside the host cell (transformation). Since, not all the cells get transformed with the recombinant/plasmid DNA, in the absence of selectable marker, it will be difficult to distinguish between transformants and non-transformant, because role of selectable marker is in the selection of transformants.
Q.8. A mixture of fragmented DNA was electrophoresed in an agarose gel. After staining the gel with ethidium bromide, no DNA bands were observed. What could be the reason?
Ans. The reasons are as follows:
(i) DNA sample that was loaded on the gel may have got contaminated with nuclease (exo-or endo-or both) and completely degraded.
(ii) Electrodes were put in opposite orientation in the gel assembly that is anode towards the wells (where DNA sample is loaded). Since DNA molecules are-negatively charged, they move towards anode and hence move out of the gel instead of moving into the matrix of gel.
(iii) Ethidium bromide was not added at all or was not added in sufficient concentration and so DNA was not visible.
Q.9. Describe the role of CaCl2 in the preparation of competent cells?
Ans. CaCl2 is known to increase the efficiency of DNA uptake to produce transformed bacterial cells. The divalent Ca2+ ions supposedly create transient pores on the bacterial cell wall by which the entry of foreign DNA is facilitated into the bacterial cells.
Q.10. What would happen when one grows a recombinant bacterium in a bioreactor but forget to add antibiotic to the medium in which the recombinant is growing?
Ans. In the absence of antibiotic, there will be no pressure on recombinants to retain the plasmid (containing the gene of your interest). Since, maintaining a high copy number of plasmids is a metabolic burden to the microbial cells, will thus tend to lose the plasmid.
Q.11. Identify and explain steps ‘A’, ‘B’ and ‘C’ in the PCR diagram given below.
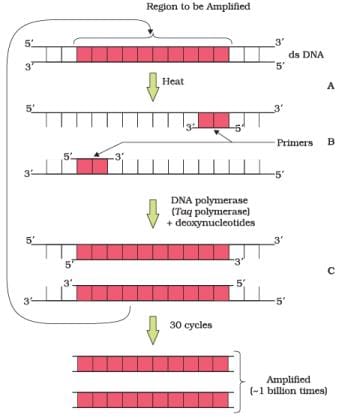
Ans.
A—Denaturation
B—Annealing
C—Extension of primers
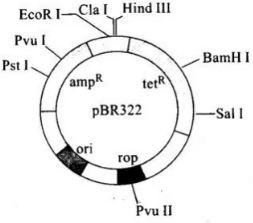
Q.12. Name the regions marked A, B and C.
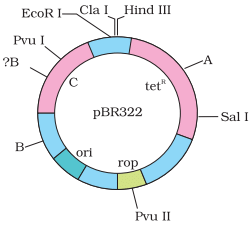
Ans.
A—Bam HI
B—Pst I .
C—Ampicillin resistance gene (ampR)
LONG ANSWER TYPE QUESTIONS
Q.1. For selection of recombinants, insertional inactivation of antibiotic marker has been superceded by insertional inactivation of a marker gene coding for a chromogenic substrate. Give reasons.
Ans. Selection of recombinants due to inactivation of antibiotics is a laborious process as it requires:
(i) a vector with two antibiotic resistance marker
(ii) preparation of two kinds of media plate, with one antibiotic each.
Transformed cells are first plated on that antibiotic plate which has not been insertionally inactivated (ampicillin) and incubated overnight for growth of transformants. For selection of recombinants, these transformants are Replica plated on second antibiotic (tetracycline) plate (which got inactivated due to insertion of gene). Non-recombinants grow on both the plates (one carrying ampicillin and the other carrying tetracycline) while recombinants will grow only on ampicillin plate.
This entire exercise is laborious and takes more time (two overnight incubation) as well. However, if we choose the second option (insertional inactivation of a marker that produces colour in the presence of a chromogenic compound), we can distinguish between the recombinants and non-recombinants on a single medium plate (containing one antibiotic and the chromogenic compound) after overnight growth.
Hence I would choose a marker which produces a coloured compound but gets inactivated due to insertion of foreign DNA.
Q.2. Describe the role of Agrobacterium tumefaciens in transforming a plant cell.
Ans. Agrobacterium tumefaciens harbours a mega plasmid called Ti-plasmid.
This has a T-DNA region flanked by left border and right border sequence. The T-DNA gets transferred and integrates with the host plant DNA. This property of Ti-plasmid has been exploited for cloning of gene of interest and stably integrating them in the plant genes. Therefore, by using Ti-plasmid or its derivatives, recombinant plant cells with desired genes of interest stably integrated in the plant genome has been successfully produced.
Q.3. Illustrate the design of a bioreactor. Highlight the difference between a flask in your laboratory and a bioreactor which allows cells to grow in a continuous culture system.
Ans. Small volume cultures cannot yield appreciable quantities of products. To produce in large quantities, the development of bioreactors, where large volumes (100-1000 litres) of culture can be processed, was required. Thus, bioreactors can be thought of as vessels in which raw materials are biologically converted into specific products, individual enzymes, etc., using microbial plant, animal or human cells. A bioreactor provides the optimal conditions for achieving the desired product by providing optimum growth conditions (temperature, pH, substrate”, salts, vitamins, oxygen).
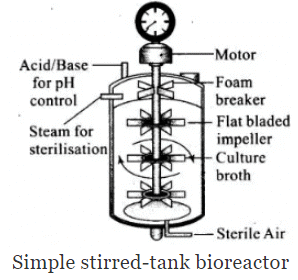
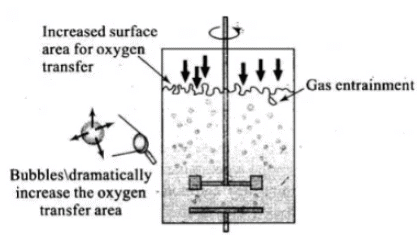
Sparged stirred-tank bioreactor through which sterile air bubbles are sparged
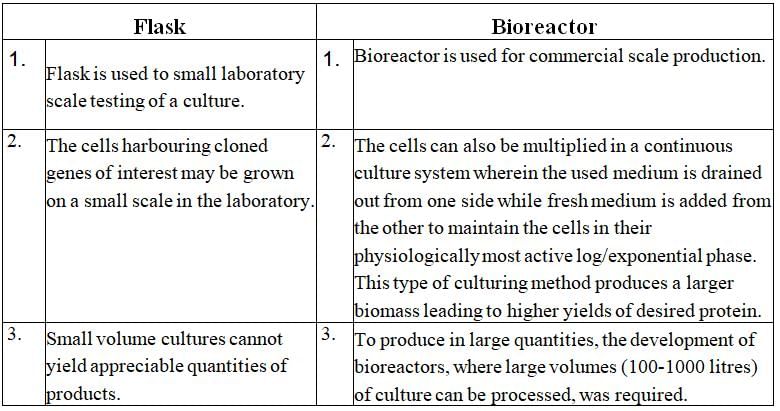
|
59 videos|290 docs|168 tests
|
FAQs on NCERT Exemplar: Biotechnology: Principle & Processes - Biology Class 12 - NEET
| 1. What are the principles of biotechnology? |  |
| 2. What are the processes involved in biotechnology? |  |
| 3. How is genetic engineering used in biotechnology? |  |
| 4. What is the significance of tissue culture in biotechnology? |  |
| 5. How is fermentation used in biotechnology? |  |
















