Genetics | Additional Study Material for NEET PDF Download
GENE EXPRESSION
The second important characteristic (first is transmission) of the gene is to store and express the genetic information that will contribute towards the phenotype.
DNA carries information for the synthesis of all proteins required for the function of a cell. A close relationship between genes and enzymes (or gene control the metabolism) was first discovered by a British physician Archibald Garrod in 1909. He observed that certain hereditary diseases in man such as alkaptonuria (black urine disease), albinism (absence of melanin pigment) phenylketonuria etc. are ‘inborn errors of metabolism and are due to the defect in the enzyme that catalyses the conversion of one metabolic substance to another. Thus, the inherited genetic defect is reflected in the deficiency of an enzyme.
Beadle and Tatum’s Hypothesis (One gene one enzyme hypothesis) The concept that genes have the information to produce enzymes, or gene metabolism relationship was experimentally proved by Beadle and Tatum (1948), on the basis of experiments conducted on pink bread mould (Neurospora crassa). This mould can normally grow in a simple minimal medium containing salts and sugar, making all other chemicals such as amino acids, purines, pyrimidines etc. through enzyme catalysed reactions. This wild type of the mould is called 'prototroph'. Beadle and Tatum exposed the pink bread mould to X-rays, which can bring about a change in the nucleotide sequence of DNA and thus causes mutation. They found that the mutants were unable to grow on a minimal medium. Each type of mutant required some extra nutrient in the minimal medium for it's normal growth. Such nutritional mutants are called 'auxotrophs'. They obtained different nutritional mutants requiring amino acids ornithine or citrulline or arginine for growth. The mutants could be classified into three types:
(i) some could grow on ornithine -, or citrulline -, or arginine - containing medium,
(ii) some could grow on citrulline - or arginine containing medium, and
(iii) some could grow only on arginine supplemented medium Effect of medium supplements on the growth of mutants Neurospora crassa
| Growth on Medium supplemented with | |||
| Mutant Type | Ornithine | Citrulline | Arginine |
I II III | + - - | + + - | + + + |
| + = growth, – = no growth. | |||
Biochemical Pathway

- This shows that all mutants could grow on arginine supplemented medium suggesting that arginine is the final product of this pathway.
- Since some mutants could not grow on ornithine, this compound must be synthesized earlier than citrulline and/ or arginine.
- Beadle and Tatum taking clue from this growth behaviour of Neurospora and other experiments, proposed the arginine biosynthetic pathway. The precursor compound first gives rise to ornithine, ornithine then gives rise to citrulline and the latter is finally converted into arginine; each of these steps is mediated by an enzyme.
- The mutant of class I, could grow on ornithine, citrulline or arginine, suggesting that they lacked the capacity to synthesize ornithine, but beyond that they could complete the pathway.
- Similarly, mutants of class II, could not convert ornithine to citrulline, but if supplied citrulline could synthesize arginine.
- The last mutants of class III could not convert citrulline to arginine and had to be supplied latter for growth.
- Beadle and Tatum reasoned that these defects could arise due to defective enzymes in each case. Since such changes were mutational, they held that one gene controls one enzyme in a pathway leading to their famous ‘one gene one enzyme hypothesis’.
The hypothesis states that each gene controls the synthesis of a specific enzyme or protein. Beadle and Tatum were awarded with Noble prize for this work in 1958. Their work founded the new science of ‘biochemical genetics’. Beadle and Tatum used total isolation method to detect the nutritional mutant of Neurospora.
Modification of Beadle and Tatum’s Hypothesis - One gene one enzyme hypothesis held that a gene has information to produce one enzyme. The enzymes are proteins, but all proteins are not enzymes. Recently some RNAs have also been found to manifest enzyme activity. Proteins are complex molecules and may be composed of one or more polypetide chains. For example, haemoglobin consists of four polypetide chains-2α and 2β chains. Thus, more than one gene may control the synthesis of a protein. It is now held that one gene is responsible for the formation of one polypetide chain. Therefore, one gene one enzyme hypothesis has been replaced by ‘one gene one polypetide hypothesis’. Further modification came when one gene was identified as a functional unit or cistron and the same was called ‘one cistron-one polypeptide hypothesis’.
TYPES OF GENE
All the genes do not play the same role nor all genes are active all the time. With regard to their role and activity, the genes are of following types:
Jumping genes – It is a segment of DNA which moves from one chromosome to another chromosome within the genome of an individual. McClintock (1983) got nobel prize for the discovery of jumping gene in maize. Two transposable controlling elements (Ds) and activator (Ac), which can jump to any chromosome from their original location on chromosome 9. Also in bacteria plasmid transposone carry gene for antibiotic resistance (ampicilline).
Other Examples : Transposable elements (TE) in Drosophila– As much as 10% of the genome consist of transposons, most important of these are copia like element, Fold back (FB) (for eye colour), and P and I element ( for sterlity). Ty -element in yeast.
- Overlapping gene – A few genes in some animal viruses code for two different polypeptides. These are called overlapping genes. For example–in φ x 174 virus, SV-40 virus.
- Constitutive genes (House-keeping genes) – These genes are expressed constantly, because their products are constant needed for cellular activity. e.g. genes for glycolysis, gene of ATPase enzyme.
- Non-constitutive genes (Smart gene or Luxary gene) – These genes remain silent and are expressed only when the gene product is needed. They are switched ‘on’ or ‘off’ according to the requirement of cellular activities. Non-constitutive genes are of two types; inducible and repressible. The inducible genes are switchedon in presence of a chemical substance called inducer, required for the functioning of gene activity. The repressible genes continue to express themselves till a chemical, often an end product of the metabolism inhibits or represses their activity. Such type of inhibition is called feedback inhibition or feedback repression.
- Homeotic genes – Homeotic gene regulates the organ differentiation in embryo. Homeobox related to transcription of homeotic gene. If mutation takes place in homoetic gene organ formation is disturbed.
- Pseudoallele – Genes lying side by side, producing related phenotypic effect distinguishable through a rare crossing over. Eg. star (dominant) and asteroid (recessive) traits inDrosophila.
- Isoallele – If several alleles exhibit same phenotype then they are said to Isoallele.
In Drosophila allele W+c W+s W+g → produce red eye colour
Hybrid vigour/Heterosis - Superiority of offsprings over it's parents is called as Hybrid vigour& it develops due to Heterozygosity.
- Hybrid vigour can be maintained for long time in vegetaively propagated crops.
- Hybrid vigour can be lost by inbreeding (selfing) because inbreeding induces the Homozygosity in offsprings. Loss of Hybrid vigour due to inbreeding, is called as inbreeding depression.
REGULATION OF GENE EXPRESSION
- The mechanism which stimulates the expression of certain genes and inhibits that of others is called regulation of gene expression.
- It is possible only if the organism has a mechanism of regulating gene activity by allowing some to function and others to restrain their activity through switching on and switching off system. This means, the genes are turned ‘on’ or ‘off’ as per requirement.
- A set of genes is ‘switched on’ when enzymes are required to metabolize a new substrate. The enzymes produced by these genes metabolize the substrate.
- The molecules of metabolite that come to switch on of the genes are termed as inducers and the phenomenon is called induction.
- Similarly, certain genes which are in their ‘switch on’ state, continue to synthesis a metabolite till the later is produced in amount more than required or else, it is supplied to the cell from outside. In other words, certain genes continue to express themselves till the end product of inhibits or repress their expression. Inhibition by an end product is known as ‘feed back repression’.
OPERON CONCEPT
- In 1961, two French microbiologist Francis Jocob and Jacques Monad at the Pasteur Institute in Paris, proposed a mechanism called operon model for the regulation of gene action in E. coli.
- An operon is a part of genetic material or DNA, which acts as a single regulated unit having one or more structural genes-an operator gene, a promoter gene, a regulator gene.
- Operons are of two types
(i) inducible
(ii) repressible.
1. Inducible System (Lac operon of E. coli) – An inducible operon system normally remains in switched off condition and begins to work only when the substance to be metabolised by it is present in the cell. Inducible operon system generally occurs in catabolic pathways.e.g.Lac operon of E. coli.
Active repressor + inducer = inactive repressor


An inducible operon system consists of four types of genes
(i) Structural genes - These genes synthesise mRNAs, which in turn synthesize polypeptide or enzyme over the ribosomes. An operon may have one or more structural genes. Each structural gene of an operon is called cistron. The lac operon (lactose operon) of Escherichia coli contains three structural genes (Z, Y and A). These genes occur adjacent to each other and thus are linked. They transcribe a polycistronic mRNA molecule (a single stretch of mRNA covering all the three genes), that helps in the synthesis of three enzymes-β galactosidase (breaks lactose into glucose and galactose), lactose permease (helps in entry of lactose in cell from outside) and transacetylase (transfers an acetyl group from acetyl Co A to β galactosidase).
(ii) Operator gene - It lies adjacent to the structural genes and directly controls the synthesis of mRNA over the structural genes. It is switched off by the presence of a repressor. An inducer can take away the repressor and switch on the gene that directs the structural genes to transcribe.
(iii) Promoter gene - This gene is the site for initial binding of RNA polymerase. When the operator gene is turned on, the enzyme RNA polymerase moves over it and reaches the structural genes to perform transcription.
(iv) Regulator gene - It produces a repressor that binds to operator gene and stops the working of the operator gene.
(v) Repressor - It is a protein, produced by the regulator gene. It binds to the operator gene so that the transcription of structural gene stops. Repressor has two allosteric site
(1) operator gene
(2) effective molecule (inducer/co-repressor)
(vi) Inducer - It is a chemical (substrate, hormone or some other metabolite) which after coming in contact with the repressor, forms an inducer repressor complex. This complex cannot bind with the operator gene, which is thus switched on. The free operator gene allows the structural gene to transcribe mRNA to synthesise the enzymes.
The inducer for lac operon of Escherichia coli is lactose (in fact allolactose an isomer of lactose). When the sugar lactose is added to the culture of E. coli,a few molecules of lactose gets into the bacterial cells by the action of the enzyme permease, a small amount of this enzyme is present in the cell even when the operon is not working. These few lactose molecules are then converted into an active form which acts as an inducer and binds to the repressor protein. The inducer repressor complex fails to join with the operator, which is turned on. The three genes are expressed as three enzymes to metabolize lactose. Allo-lactose is real inducer of lac operon.
Gratuitous inducer → Some molecules resemble with natural inducer but are not metabolize by the enzyme.
Example :Isopropyl thio galactoside (IPTG): This resembles lactose and thus has the property of induction.
Such inducer which induces enzyme synthesis, without getting metabolized are called gratuitous inducer.
2. Repressible System (Tryptophan operon of E. coli.) – A repressible operon system is normally in it's switch on state and continue to synthesize a metabolise till the latter is produced in amount more than required, or else it becomes available to the cell from outside. Repressible operon system is commonly found in anabolic pathway.
e.g. Tryptophan operon of E. coli. [Inactive repressor + co-repressor = acitve repressor] Tryptophan operon of Escherichia coli is an example of repressible system. It consists of the following:
(i) Structural genes. These genes are meant for transcription of mRNA, which in turn synthesize enzyme.
Tryptophan operon has five structural genes E, D, C, B and A. They lie in continuation and synthesize enzymes for five steps of tryptophan synthesis.
(ii) Operater gene (trp O). It lies adjacent to the structural genes and controls the functioning of the structural genes. Normally, it is kept switched on, because the apo-repressor produced by the regulator gene does not bind to it. The operator gene is switched off when a co-repressor is available along with apo-repressor.
(iii) Promoter gene (trp P). It marks the site at which the RNA polymerase enzyme binds. When the operator gene is switched on, it moves from promotor gene to structural genes for transcription.
(iv) Regulator gene (trp R). It produces a regulatory protein called apo-repressor for (Inactive repressor) possible blocking the activity of operator gene.
(v) Apo-repressor. It is a regulatory protein synthesised by regulator gene. When a co-repressor substrate is available in the cell, the apo-repressor combines with the co-repressor to form a apo-repressor co-repressor complex. This complex binds with the operator gene and switches it off. Presence of aporepressor alone, the operator gene is kept switched on because, by itself the apo-repressor is unable to block the working of operator gene.
(vi) Co-repressor. It is an end product of reactions catalysed by enzymes produced by the structural genes.
In the presence of tryptophan some molecules of tryptophan act as co-repressor, co-repressor-bind with inactive repressor. co-repressor repressor complex bind with-operator region and prevent the binding of RNA polymerase to the promoter, the trp-operon is off.


| Differences between Induction and Repression | |
| Induction | Repression |
1. It is switching-on of an operon which is normally in switched-off state. 2. It starts transcription and translation. 3. It is caused by a new metabolite which needs enzymes to get metabolised. 4. It generally operates in a catabolic pathway. | 1. It is switching-off an operon which is normally in switched-on state: 2. It stops transcription and translation. 3. It is caused by increased formation or availability of a metabolite. 4. It generally operates in an anabolic pathway. |
The repressor molecule has key role in regulation of lac-operon. Repressor molecule active or inactive.
Active repressor may be rendered inactive by addition of an inducer while the inactive repressor can be made active by addition of a co-repressor.
Because the product of regulator gene the repressor act by shutling off the transcription of structural gene the operon model, as originally proposed by Jocob & Monad is referred as –negative control system.
GENE EXPRESSION IN EUKARYOTES
1. In eukaryotes, functionally related genes may not be clustered together constituting an operon.
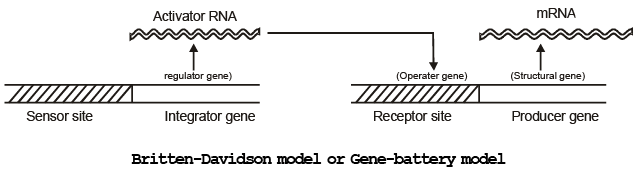
2. The most popular model is known as ‘Britten-Davidson model’ or ‘Gene-battery model’ proposed by Britten and Davidson in 1969.
3. A set of structural genes controlled by one sensor site is termed as battery.
4. Gene-battery model assumes the presence of four classes of sequences:
(i) Producer gene: A producer gene is comparable to structural gene of prokaryotic operon.
(ii) Receptor site: A receptor site is comparable to operater gene of bacterial operon and one such receptor site is assumed to be present adjacent to each producer gene.
(iii) Integrator gene: Integrator gene is comparable to regulator gene and is responsible for synthesis of an activator RNA. It activates the receptor site.
(iv) Sensor site: A sensor site regulates the activity of integrator gene. Activator gene can be transcribed only when the sensor site is activated.
The sensor sites are recognized by agents which change the patterns of gene expression like hormones and proteins. When a transcription factor (protein, hormone) bind to the sensor site it cause the transcription of integrator.
HUMAN GENETICS
The study (analysis) of genetic characters and aspects like genetic improvements among humans are included in human genetics. This is also known as eugenics (= well born).
Eugenics is a term derived from Greek language 'Eugenes' meaning "Well born". First of all, Sir Francis Galton, 1883 proposed the idea of improvement in human species through change in hereditary characters in a scientific manner and named it Eugenics. Because of this Sir Francis Galton is known as "father of Eugenics".
To find out various facts, scientists have to perform a number of experiments, but it is not possible to do so in humans. The following problems are faced in studying human genetics:-
1. Cells of human body are relatively smaller and number of chromosomes present in them is more.
2. It is not possible to perform various experiments on humans in the laboratory.
3. Due to greater life span of humans, a lot of time is required to study their genetic characteristics.
4. Rate of reproduction is slow in humans.
5. Individuals of human species are generally heterozygous for various characters and it is very difficult to find homozygous individuals.
6. Due to controlled hybridization and long lived life among humans, it is not possible to study many generations in easy way.
Despite of above mentioned problems humans are considered suitable for genetical experiments due to
1. Longer life span of humans, the abnormalities that require relatively long period to express themselves can be easily studied.
2. By pedigree study and analysis of families, many genetic characters of man can be traced out.
3. To study many diseases (like haemophilia, colour blindness etc.) and intelligence quotient (I.Q.) humans are more preferable than any other organisms.
Examples of some autosomal characters in human
Character | Dominant | Recessive |
Eye colour | Brown / Black | Blue |
Eye size | Large | Small |
Hair | Curly hair | Straight |
Cheek | Dimple cheek | Normal |
Hand | Right Handed | Left handed |
Rolling of tongue | Rollar | Non Rollar |
Rh Factor | Rh+ | Rh — |
PTC taster | Taster | Non Taster |
Skin pigmentation | Normal | Albino |
Devices Used in Human Genetical Studies The study and analysis of human genetics is performed by many methods like pedigree analysis, statistical analysis and human karyotyping. Of these the important ones, that is, pedigree analysis and human karyotype is being described here.
1. Pedigree Analysis Study of ancestoral history of man of transmission of genetic characters from one generation to next, is pedigree analysis. Dwarfism, albinism, colour blindness, haemophilia etc. are genetically transmitted characters. To study and analyse them a pedigree of genetic facts/data and following symbols are used.
Symbol used in Pedigree :
1. – Normal Male
– Normal Male
2.  – Normal Female
– Normal Female
3.  – Mating (marriage) Usual practice is to place the males first, on the left if one parent is not shown in a pedigree, it indicates that this individual was phenotypically normal.
– Mating (marriage) Usual practice is to place the males first, on the left if one parent is not shown in a pedigree, it indicates that this individual was phenotypically normal.
4.  The siblings are indicated in chronological order of birth
The siblings are indicated in chronological order of birth
5.  Sex unspecified
Sex unspecified
6. Twin If monozygotic
If dizygotic 
7.  , Affected male and female individual
, Affected male and female individual
8.  , Heterozygous for autosomal recessive
, Heterozygous for autosomal recessive
9.  Carrier female of sex linked recessive character or disease
Carrier female of sex linked recessive character or disease
10.  Death of individual
Death of individual
11.  Abortion or still birth (sex unspecified)
Abortion or still birth (sex unspecified)
12.  Consanguineous marriage
Consanguineous marriage
13.  Parent with male child affected with disease
Parent with male child affected with disease
14.  – Five unaffected offsprings
– Five unaffected offsprings
Pedigree analysis provides valuable informations regarding genetical make up of human beings. If any genetic disease is occurring in a family, then pedigree analysis provides guidance to forthcoming parents about their future progenies for example- polydactyly in humans.
2. Human Karyotype Humans have 23 pairs (46) chromosomes. In this method, the chromosomes (autosomes and sex chromosomes) are arranged according to their size and structure. Based on the position of centromere and relative lengths of both arms of chromosome, three types of chromosomes are found in human - metacentric, submetacentric and acrocentire. Karyotype helps to know the relative structures (morphology) of chromosomes. Besides, it helps in chromosomal identification and it's nomenclature. It is also used in studying chromosomal abnormalities.
Eugenics For improvement of hereditary characters in future generations there should be an increase in number of best individuals and reduction in number of defective individuals. Presently, eugenics is studied under following types -
1. Positive Eugenics In positive eugenics the main approach is to increase the number of progenies having best hereditary characters.
This is achieved through genetic counselling, selective mating (planned marriages), arranging marriages at right age, freedom from social bonds etc.
Preservation of ovum of superior quality, artificial insemination and preserving the best quality sperms with aid of modern techniques are also included in positive eugenics.
Euphenics and gene engineering techniques are also being used to establish the best quality of hereditary traits in human society.
2. Negative Eugenics In negative eugenics the carriers of undesirable inferior traits (or characters or genes) are not allowed to produce their progenies by various checks such as marriage control, segregation, birth control, sterilization and control of consanguineous marriages.
Example : Given pedigree shows inheritance of albinism which is an autosomal recessive disorder. If 4th individual is homozygous normal then find out the carrier individual.


Ans. : 1, 2, 5, 6 (Aa)
POPULATION GENETICS
Study of gene frequency in a population is called population genetics.
1. Gene pool – A gene pool is the sum total of genes in reproductive gametes of a population.
2. Gene-flow – Migration of gene from one population to another population by cross fertilization.
3. Genetic load – The existence within the population of dis-advantegeous allele in heterozygous genotype is known as genetic load
4. Gene frequency – Gene frequency is defined as proportion of different alleles of a gene in a population.
Ex. In a population of 100 individuals of MN blood group 50 MM, 20 MN, 30 NN find out the frequency of
M & N.
M M – 50
M N – 20
N N – 30
Total M gene – 50 × 2 + 20 = 120
Total N gene – 30 × 2 + 20 = 80
Total gene – 200
Freq. of M gene

Freq. of N gene

0.6 + 0.4 = 1
p + q = 1
Hardy Weinberg Law – 1908 G.H.Hardy (English mathematician) & German Physician, W.Weinbergh independently discovered that an equilibrium is established between frequencies of allele in random mating population and these gene frequency remain constant from generation to generation.
This law applicable when factors like mutation, selection & migration, are absent.
Hardy Weinberg theorem or Hardy weinbergh law
nberg theorem or Hardy weinbergh law
p2 + 2pq + q2 = 1 (A → p, a → q, p+q = 1)
AA Aa aa
In this equation frequency of A → P or the Frequency of Homozygous dominant will be AA–P2 In this equation frequency of a → q the frequency of homozygous recessive – q2 the frequency of Aa = 2pq The frequency of different genotype produced due to random mating will depend upon the gene frequency and equilibrium is established after one single generation of random mating.
Factors affecting gene frequency :
(1) Mutation
(2) Selection
(3) Migration
(4) Random drift
Directional-mutation, selection, migration Non-directional = random drift
Non-directional which cannot be predicted from one generation to the other.
1. Mutation : Mutation lead to introduction of new gene leading to genetic difference. Change in gene frequency due to mutation, will depend upon mutation rate.
2. Selection (Natural selection) : By the natural selection there is increase in frequency that are more fit to nature.
Selections are associated with superior fitness will increase in frequency in the population.
3. Migration : Migration recessive or dominant allele can be introduced into a population from a nearby population.
This causes difference in frequency between the two population.
4. Random drift or Genetic drift.: Allele frequency can change randomly, sometime increasing and sometime decreasing such deviation in gene frequency will be due to sampling error.
Due to sampling error, random changes in gene frequencies are called random drift.

σ – sampling error
p & q – initial gene frequency
N – No. of genes which are sampled
population  sampling error
sampling error 
population  sampling error
sampling error 
Q. In a random population frequency of recessive phenotype is 0.09. What is the frequency of heterozygous genotype?
Sol. q2 = 0.09
q = 0.3
p = 1 – q
p = 1 – 0.3 ⇒ 0.7
Frequency of heterozygote = 2pq = 2 × 0.7 × 0.3 = 0.42 = 42%
GENETIC ENGINEERING
INTRODUCTION :
- Genetic engineering also referred as ‘recombinant DNA technology’ or ‘gene splicing’ is one kind of biotechnology involving manipulation of DNA.
- It deals with the isolation of useful genes from a variety of sources and the formation of new combinations of DNA (recombinant DNA) for repair, improvement, perfection and matching of a genotype.
- Thus, genetic engineering may be defined ‘as a technique for artificial and deliberately modifying DNA (gene) to suit human needs’.
- In genetic engineering breakage of DNA molecule at two desired places is done with the help of restriction endonuclease to isolate a specific DNA segment and then insert it in another DNA molecule at a desired position.
- The new DNA molecule is recombinant DNA and the technique called genetic engineering. Genetic engineering aims at adding, removing or repairing of a part of genetic material. Genetic engineering can be used to improve the quality of human life.
Paul bergh (Father of genetic engineering). He transferred gene of SV-40 virus (simian virus) in to E.coli with the help of λ – phage. (Nobel prize - 1980) The concept of genetic engineering was the outcome of two very significant discoveries made in bacterial research. These were–
(i) presence of extra-chromosomal DNA fragments called plasmids in the bacterial cell,which replicate along with chromosomal DNA of the bacterium.
(ii) presence of enzymes restriction endonucleases which cut DNA at specific sites. These enzymes are, therefore, called ‘molecular scissors’.
TOOLS AND TECHNIQUES OF GENETIC ENGINEERING
Tools
Genetic engineering involves cutting of desired segments of DNA and pasting of this D.N.A in a vector to produce a recombinant DNA (rDNA). The ‘biological tools’ used in the synthesis of recombinant DNA include enzymes, vehicle or vector DNA, passenger DNA and alkaline phosphatases.
1. Enzymes. A number of specific kinds of enzymes are employed in genetic engineering.
These include lysing enzymes, cleaving enzymes, synthesising enzymes and joining enzymes.
(i) Lysing enzymes. These enzymes are used for opening the cells to get DNA for genetic experiment.
Bacterial cell wall is commonly dissolved with the help of lysozyme.
(ii) Cleaving enzymes. These enzymes are used for DNA molecules. Cleaving enzymes are of three types; exonucleases, endonucleases and restriction endonucleases.

(a) Exonucleases cut off nucleotides from 5' or 3' ends of DNA molecule.
(b) Endonucleases break DNA duplex at any point except the end.
(c) Restriction endonucleases cleave DNA duplex at specific points in such a way that they come to possess short single stranded free ends.
For example, a restriction endonuclease ECOR-l (from Escherichia coli) recognizes the base sequence GAATTC/CTTAAG in DNA duplex and cleaves it's strands between G and A.
Restriction enzymes are obtained from bacteria. They are useful to bacteria because the enzyme bring about fragmentation of viral DNA without affecting the bacterial genome. This is an adaptation against bacteriophages.
Restriction enzyme (Eco R-I) was discovered by Arber, Smith & Nathans (1978 Nobel prize). These enzymes exist in many bacteria beside cleavage some restriction endonuclease, also have capability of modification.
Modification in the form of methylation, by methylation the bacterial DNA modifies and therefore protects it's own chromosomal DNA from cleavage by these restriction enzymes.
Restriction enzymes are used in recombinant DNA technology because they can be used in vitro to recognize and cleave within specific DNA sequence typically consisting of 4 to 8 nucleotides. This specific 4 to 8 nucleotide sequence is called restriction site and is usually palindromic,this means that the DNA sequence is the same when read in a 5'-3' direction on both DNA strand

AND MADAM DNA
As a result the DNA fragments produced by cleavage with these enzymes have short single stranded overhang at each end these kinds of ends are called sticky or cohesive ends because base pairing between them can stick the DNA molecule back together again.

Therefore by cutting two different DNA samples with the same restriction enzyme and mixing the fragments together a recombinant DNA molecule can be generated.
Exceptionally, some enzymes cleave both strand of DNA at exactly the same nucleotide position, typically in the center of the recognition sequence resulting in blunt end or flush end.
Sma I (Serratia marcescens)

Nomenclature of enzyme –The first letter used for the enzyme is the first letter of the bacterium genus name (in Italics) then comes the first two letter of it's species (In Italics), next is the strain of the organism, last is Roman numerical signifying the order in which the enzymes were isolated from that strain of bacteria.
EXAMPLES OF RESTRICTION ENZYME
Recognition sequences of some restriction endonucleases
| Name | Recognition sequence | End after cleavage | Source | |
Eco RI | | – G | A A T T C – | Escherichia coli - containing drug resistant plasmid RI. |
| Hind III |  | – A – T T C G A | A G C T T – A – | Haemophilus influenzae |
| Bam I |  | – G – C C T A G | G A T C C – G – | Bacillus amyloliquefaciens |
| Hae III |  | – G G – C C | C C – G G – | Haemophilus aegyptius |
(iii) Synthesizing enzymes. These enzymes are used to synthesize new strands of DNA, complementary to existing DNA or RNA template. They are of two types; reverse transcriptases and DNA polymerases.
(a) Reverse transcriptases help in the synthesis of complementary DNA strands on RNA templates;
(b) DNA polymerases help in the synthesis of complementary DNA strands on DNA templates.
(iv) Joining enzymes. These enzymes help in joining the DNA fragments. For example DNA ligase from Escherichia coli is used to join DNA fragments. Joining enzymes are, therefore, called molecular glues.
(iv) Alkaline phosphatases. These enzymes cut off phosphate group from the 5' end of linearised circular DNA and prevent its recircularization.
2. Vehicle DNA or Vector DNA. The DNA used as a carrier for transferring a fragment of foreign DNA into a suitable host is called vehicle or vector DNA.
You know that plasmids and bacteriophages have the ability to replicate within bacterial cells independent of the control of chromosomal DNA.
The following are the features that are required to facilitate cloning into a vector.
(i) Origin of replication (ori) : This is a sequence from where replication starts and any piece of DNA when linked to this sequence can be made to replicate within the host cells. This sequence is also responsible for controlling the copy number of the linked DNA. So, if one wants to recover many copies of the target DNA it should be cloned in a vector whose origin support high copy number.
(ii) Selectable marker : In addition to 'ori', the vector requires a selectable marker. Normally, the genes encoding resistance to antibiotics such as ampicilin, chloramphenicol, tetracycline or kanamycin, etc., are considered useful selectable markers for E. coli.

(iii) cloning sites : In order to link the alien DNA, the vector needs, recognition sites for the commonly used restriction enzymes. The ligation of alien DNA is carried out at a restriction site present in one of the two antibiotic resistance genes. For example, you can ligate a foreign DNA at the Bam H I site of tetracycline resistance gene in the vector pBR322. The recombinant plasmids will loss tetracycline resistance due to insertion of foreign DNA (insertional inactivation) now, it can be selected out from non-recombinant ones by plating the transformants on ampicillin containing medium. The transformants (plasmid transfer) growing on ampicillin containing medium are then transferred on a medium containing tetracycline. The recombinants will grow in ampicillin containing medium but not on that containing tetracycline. But, nonrecombinants will grow on the medium containing both the antibiotics. In this case, one antibiotic resistance gene helps in selecting the transformants.
Selection of recombinants due to inactivation of antibiotics is a cumbersome (troublesome) procedure because it requires simultaneous plating on two plated having different antibiotics. Therefore, alternative selectable markers have been developed which differentiate recombinants from non-recombinants on the basis of their ability to produce colour in the presence of a chromogenic substrate. In this, a recombinant DNA is inserted within the coding sequence of an enzyme, which is referred to as insertional inactivation.
The presence of a chromogenic substrate X–gal (5–bromo–4chloro–β-D galacto pyranoside) gives blue coloured colonies if the plasmid in the bacteria does not have an insert. Presence of insert results into insertional inactivation of the b-galactosidase (reporter enzyme) and the colonies do not produce any colour, these are identified as recombinant colonies.
(iv) Vectors for cloning genes in plants and animals : Agrobacterium tumifaciens, a pathogen of several dicot plants deliver a piece of DNA known as 'T-DNA' to transform normal plant cells into a tumour .
Similarly, retroviruses in animals have the ability to transform normal cells into cancerous cells. A better understanding of the art of delivering genes by pathogens in their eucaryotic hosts has generated knowledge to transform these tools of pathogens into useful vectors for delivering genes of interest to humans. The tumour inducing (Ti) plasmid of Agro-bacterium tumifaciens has now been modified into a cloning vector which is no more pathogenic to the plants but is still able to use the mechanisms to deliver genes (disarmed) and are now used to deliver desirable genes into animal cells. So, once a gene or a DNA fragment has been ligated into a suitable vector it is transferred into a bacterial, plant or animal host (where it multiplies).
(i) Plasmids. They are extra chromosomal DNA segments found in bacteria which can replicate independently. Plasmids can be taken out of bacteria and made to combine with desired DNA segments by means of restriction enzymes and DNA ligase. A plasmid carrying DNA of another organism integrated with it, is known as recombinant plasmid or hybrid plasmid or Chimeric plasmid.
(ii) Viruses. The DNA of certain viruses is also suitable for use as a vehicle DNA. Bacteriophage (bacterial virus) has been used to transfer gene for b galactosidase from Escherichia coli to human cells. Lambda phage (l phage) has been used for transferring lac genes of Escherichia coli into haploid callus of tomato.
Vector type | Insert size kb |
Plasmid | 0.5 – 8 |
Bacterophage lambda | 9 – 23 |
Cosmid | 30 – 45 |
BAC | 50 – 300 |
YAC | 1000 – 2500 |

3. Passenger DNA. It is the DNA which is transferred from one organism into another by combining it with the vehicle DNA. The passenger DNA can be complementary, synthetic or random.
(i) Complementary DNA (cDNA)- It is synthesized on mRNA template with the help of reverse transcriptase and necessary nucleotides. The DNA strand is then separated from the hybrid RNA-DNA complex by using alkali. Complementary DNA strand is then synthesized over the template of cDNA with the help of DNA polymerase. cDNA formed through reverse transcription is shorter than the actual or in vivo gene because of the absence of introns or non-coding regions.
(ii) Synthetic DNA (sDNA)- It is synthesized with the help of DNA polymerase on DNA template.
Kornberg (1961) synthesized first synthetic DNA from a mixture of deoxyribo nucleotide triphosphates, DNA polymerase enzyme, metal ions and a segment of viral DNA.
Khorana (1968) synthesized first artificial gene (DNA) without a template. They synthesized the gene coding for yeast alaninet-RNA,which contained only 77 base pairs. However, it did not function in the living system. In 1979, Khorana was able to synthesize a functional tyrosine t-RNA gene of E. coli with 207 nucleotide pairs. Since then a number of genes have been synthesized artificially.
(iii) Random DNA - It refers to small fragments formed by breaking a chromosome with the help of restriction endonucleases.
Technique of Recombinant DNA
- The DNAs which are to be used as passenger DNA and the vehicle DNA are extracted out of their cells by lysing the cells with the suitable enzyme.
- The extracted DNAs are isolated from other cell contents by ultra centrifugation and purified.
- Both the passenger and vehicle DNAs are then, cleaved by using the same restriction endonuclease so that they have complementary sticky ends.
- The complementary sticky ends of the passenger and vehicle DNAs are joined with ligase enzyme. This gives rise to a recombinant DNA.
- The recombinant DNA is now inserted into a host cell such as Escherichia coli. The bacteria to be used as hosts should be without plasmids.
- The host cells are treated with calcium chloride or lysozyme. It creates transient (temporary) pores in their wall and makes the latter permeable to recombinant DNA.
- The recombinant DNA is added to the culture in which such bacteria are growing. The recombinant DNA is taken up by the bacteria. It replicates when the host bacteria divide and give rise to multiple copies of recombinant DNA.

Fig.: Mechanism of transferring insulin gene from human DNA to E.coli with a plasmid
GENE TRANSFER
Transfer of desired genes from one organism into another is an important aspect of genetic engineering. Gene transfer involves essentially three stages:
(i) Isolating a useful DNA segment from the donor organism.
(ii) Inserting it to a suitable vector.
(iii) Splicing of this altered DNAs into a recipient organism or host cell.
The cell to begin making the appropriate product and then device an economical method for it's mass production.
Gene transfer is achieved by two kinds of transfer methods:
(i) Indirect method through vectors or carriers and
(ii) Direct or vector less transfer method.
(i) Gene Transfer Through Vectors. Plasmids and viruses are commonly used vectors for the transfer of desired genes. The desired genes are first made to join suitable plasmid or virus which are then introduced into the target cells.

- A plant pathogenic bacterium-Agrobacterium tumefaciens produces crown galls or plant tumours in almost all dicotyledonous plants.
- This bacterium infects all broad leaved agricultural crops such as tomato, soyabean, sunflower and cotton but not cereales.
- Tumour formation is induced by it's plasmid which is therefore called Ti plasmid (Ti for tumour inducing) Agrobacterium tumefaciens naturally transfers some part of Ti-plasmid in to host plant DNA without any human effort so it is called natural genetic engineer of plant.
In the transformation process two essential component in Ti-plasmid –
(a) T-DNA – (Transferred DNA)
(b) Vir-region – (Virulence region)
Inside the host plant cell T-DNA is separated from Ti-plasmid, and integrated into host plant DNA that causes crown gall tumour.
Vir-region contains genes which are essential for T-DNA transfer and integration to host plant DNA. Vir-region contains 6-operons A, B, C, D, E, and G. (Acetosyringone is inducer of Vir-operon) When we use Ti plasmid as a vector, first we remove the tumour causing gene from T-DNA region. Then desired gene inserted inplace of it. Now, this plasmid is called disarmed plasmid.
Same as Ri plasmid of a rhizogenes (causing hairy root disease) also used as vector for gene transfer to plant cell.
(ii) Vectorless Gene Transfer : Foreign genes can also be transferred directly by the following methods:
(a) Electroporation- It creates transients (temporary pores) in the plasma membrane to facilitate entry of foreign DNA.
(b) Chemical mediated genetic transformation- It involves certain chemicals such as polyethylene glycol (PEG), that help in the uptake of foreign DNA into host cells.
(c) Microinjection- It is the introduction of foreign genes into plant or animal cells using micropipettes or glass needles.
(d) Particle gun/Biolistic method- It is a technique in which tungsten or gold particles coated with foreign DNA are bombarded into target cells to facilitate entry of the foreign genes.
(e) Liposome mediatd gene transfer- In this method DNA encloses within lipid begs. These lipid begs fused with protoplast.
GENE TRANSFER IN ANIMALS
In animals, the genes are transferred mostly through direct methods such as electroporation, microinjection or using particle gun.
The desired foreign genes can be introduced into fertilised eggs or embryos through micro-injection. These transgenic eggs or embryos can be implanted into the uterus of another female, called surrogate (foster) mother for their further development. Now, since most of the human genes have been identified through ‘human genome project’, it is hoped that a number of human genetic disorders such as Alzheimer, cancer, haemophilia, thalassemia and cystic fibrosis can now be cured through insertion of the correct genes into these patients.
ACHIEVEMENTS OF GENETIC ENGINEERING
- The DNA recombinant technology or genetic engineering provides great benefits for advancement of science and society.
(1) Gene Therapy :
- A new system of medicine gene therapy, may develop to treat hereditary diseases such as haemophilia.
- It is the technique for curing genetic disease through replacing a "Faulty Gene" by a normal healthy functional gene.
- The first gene therapy used in severe combined immuno deficiency (SCID) pateint.
About the 25% of infant with SCID disorder lack the enzyme adenosine deaminase (ADA) - ADA enzyme involved in purine metabolism.
- These patients have no functioning T & B lymphocytes.
- The affected child of SCID develops recurrent infection early in life because they have no immune response against invading pathogen.
- The ideal approach would be to give the patient a functioning ADA by gene therapy.
- Thus, genetic disorder can be over come by introducing specific gene.
(2) Bacteria may be used as "Living factories" for synthesizing vitamins, hormones and antibodies.
- Human insulin (Humulin) was first genetically engineered product produced by an American firm Eli Lilly – 5th July 1983.
- Charles weismann of university of Zurich, obtained interferon through recombinant E.coli (1980)
- Microbes have been engineered to produce Human growth Hormone (HGH) for curing dwarfism.
- Vaccines which are produced by genetic engineering e.g., for Hepatitis-B and Herpes virus).
- Nitrogen fixation genes may be transferred from bacteria to the major food crops to boost food production without using expensive fertilizers.
- Transgenic plant obtained through recombinant DNA technology. First transgenic plant was tobacco. It contains resistant gene against weedicide (Glycophosate).
First transgenic animal was mouse contain gene for growth hormone.
First introduced transgenic crop in India (2002) isBt-cotton.
It is resistant for boll worm(Helicoperpa armigera - Larva of insect). It is formed by transfer of pest resistant gene from Bacillus thuringiensis (bt-2 gene encoding Bt–toxin).
Bacillus thuringiensis produces a toxic protein called crystal protein (Cry-Protein) this protein is toxic for larva of certain insect.
This protein kills the insect by inhibiting ion transport in midgut (bt 2 gene is called cry-gene)
- In pollution control, microbes have been engineered to break up the crude oil spills.
Dr. Ananda Mohan chakraborti introduced plasmid from different strains in to single cell of pseudomonas putida. The result was new genetically engineered bacterium which would cleaning oil spills called "Super bug" (oil eating bug.)
(3) Medical Diagnosis of Disease Recombinant DNA technology is one of the important tool for the diagnosis of several diseases. The diagnostic technique involves the construction of probes consisting of short segments of single stranded DNA attached to a radioactive or fluoroscent marker. Such probes are used to identify infections agents such as Salmonella (cause food poisoning), Staphylococcus (pus forming bacterium), hepatitis virus, HIV; muscular dystrophy, cystic fibrosis and so on. Recombinant DNA technology can also be employed to predict the inheritance of genetic disorders from carrier parents. The chances of birth of an affected child can be predicted by testing DNA of repetitive prospective genetic disorder carrier parents.
Applications of Recombinant DNA products
Medically Useful recombinant products
- Human insulin
- Human growth hormone
- Calcitonin
- Chorionic gonadotropin
- Blood clotting factor VIII/IX
- Tissue Plasminogen activator (TPA)
- Erythropoetin
- Platelet derived growth factor
- Interferon
- Interleukin
- Vaccines
Applications
- Treatment of insulin - dependant diabetes
- Replacement of missing hormone in short stature people.
- Treatment of rickets.
- Treatment of infertility.
- Replacement of clotting factor missing in patients with Haemophilla A/B.
- Dissolving of blood clots after heart attacks and strokes.
- Stimulation of the formation of erythrocytes (RBCs) for patients suffering from anaemia during dialysis or side effects of AIDS patients treated by drugs.
- Stimulation of wound healing
- Treatment of pathogenic viral infections, cancer
- Enhancement of action of immune system
- Prevention of infectious diseases such as hepatitis B, herpes, influenza, pertusis, meningitis, etc.
Application of Genetically Engineered Microbes
Microbes | Applications |
Escherichia coli (gut bacterium) | Production of human insulin, human growth factor interferons, interleukin and so on. |
Bacillus thuringiensis (soil bacterium) | Productions of endotoxin (Bt toxin), highly potent, safe and biodegradable insecticide for plant protection. |
Rhizobium meliloti (bacterium) | Nitrogen fixation by incorporating "nif" gene in cereal crops. |
Pseudomonas putida (bacterium) | Scavenging of oil spills by digesting hydrocarbons of crude oil. |
Bacterial strains capable of accumulating heavy metal Trichoderma (fungus) | Bioremediation (cleaning of pollutants in the environment). |
Transgenics and their potential applications
Transgenic | Useful applications |
Bt Cotton | Pest resistance, herbicide tolerance, and high yield. |
Flavr Savr Tomato | Increased shelf-life (delayed ripening) and better nutrient quality |
Golden Rice | Vitamin A and Fe - rich |
Cattles (cow, sheep, goat) | Therapeutic human proteins in their milk |
Pig | Organ transplantation without risk of rejection |
POLYMERASE CHAIN REACTION TECHNOLOGY (PCR-TECHNOLOGY)
- This technique was invented by Kary mullis (1983).
- In 1993 Karry Mullis got nobel prize for PCR(for chemistry)
- PCR is a method for amplifying a specific region of DNA molecule without the requirement for time consuming clonning procedures.
- PCR reaction takes place in Eppendrof tube.
- Using PCR-technique very low content of DNA available from sampels of blood or semen or any other tissue or hair cell can be amplified many times and analysed. In this technique Taq-Polymerase is used. Taq polymerare enzyme is used in PCR which is a special type of DNA polymerare enzyme which is resistant to high temperature.
- Taq Polymerase is isolated from Thermus aquaticus bacterium.
- Some other examples of polymerase which are used in PCR are
- Pflu Polymerase - Isolated from Pyrococus furiosus bacterium.
Vent Polymerase - Isolated from Thermococcus litoralis bacterium.
3 Main steps in PCR :-
(1) Denaturation
(2) Annealing
(3) Extension
Denaturation (94°C)

Annealing (54°C)

Extension (72°C)

1. Denaturation :- In this step a double stranded DNA molecule is placed at 94°C. So double stranded DNA becomes single stranded & each single stranded DNA functions as a template.
2. Annealing :- In this step two primer DNA are attached at 3' end of single stranded DNA.
3. Extension :- In this process taq polymerase enzyme synthesize DNA strain over template. PCR is automatic process because taq. polymerase enzyme is heat resistant.
DNA FINGER PRINTING / DNA TYPING / DNA PROFILING/ DNA TEST
- It is technique to identify a person on the basis of his/her DNA specificity.
- This technique was invented by sir Alec. Jeffery(1984). In India DNA Finger printing has been started by Dr. V.K. Kashyap & Dr. Lal Ji Singh.
- DNA of human is almost the same for all individuals but very small amount that differs from person to person that forensic scientists analyze to identify people.
These differences are called Polymorphism (many forms) and are the key of DNA typing. Polymorphisms are most useful to forensic scientist. It is consist of variation in the length of DNA at specific loci is called Restricted fragment. It is most important segment for DNA test made up of short repetitive nucleotide sequences, these are called VNTRs (variable number of tandem repeat).
- VNTR's also called mini-satellites were discovered by Alec Jeffery. Restricted fragment consist of hypervariable repeat region of DNA having a basic repeat sequence of 11-60 bp and flanked on both sites by restriction site. The number and position of mini-satellites or VNTR in restriction fragment is different for each DNA and length of restricted fragment is depend on number of VNTR.
- Therefore, when the genome of two people are cut using the same restriction enzyme the length of fragments obtained is different for both the people.
- These variations in length of restricted fragment is called RFLP or Restriction fragment length polymorphism.
- Restriction Fragment Length Polymorphism distributed throughout human genomes are useful for DNA Finger printing.
- DNA Fingerprint can be prepared from extremely minute amount of blood, semen, hair bulb or any other cell of the body.
DNA content of 1 - Microgram is sufficient.
Technique of DNA Finger printing involves the following major steps.
1. Extraction – DNA extracted from the cell by cell lysis. If the content of DNA is limited then DNA can be amplified by Polymerase chain reaction (PCR). This process is amplification.

Steps of DNA Fingerprinting
2. Restriction Enzyme Digestion : Restriction enzyme cuts DNA at specific 4 or 6 base pair sequences called restriction site.
Hae III (Haemophilus aegyptius) is most commonly used enzyme. It cuts the DNA, every where the bases are arranged in the sequence GGCC. These restricted fragment transferred to Agarose Polymer gel.
3. Gel Electrophoresis : – Gel electrophoresis is a method that separates macromolecules-either nucleic acid or proteins-on the basis of size, electric charge.
- A gel is a colloid in a solid form. The term electrophoresis describes the migration of charged particles under the influence of an electric field. Electro refers to the energy of electricity. Phoresis, from the Greek verb phoros, means "to carry across." Thus, gel electrophoresis refers to the technique in which molecules are forced across a span of gel, motivated by an electrical current. Activated electrodes at either end of the gel provides the driving force. A molecule's properties determine, how rapidly an electric field can move the molecule through a gelatinous medium.
- Many important biological molecules such as amino acids, peptides, proteins, nucleotides, and nucleic acids posses ionisable groups and, therefore, at any given pH, exist in solution as electrically charged species either as cation (+) or anions (–). Depending on the nature of the net charge, the charged particles will migrate either to the cathode or to the anode. – By the gel electrophoresis these restricted fragments move towards the positive electrode (anode) because DNA has –ve electric charge (PO4–3).
- Smaller Fragment more move towards the positive pole due to less molecular weight. So after the gel electrophoresis DNA fragment arranged according to molecular weight
- These separated fragments can be visualized by staining them with a dye that fluoresces ultraviolet radiation.
- This appears the specific restricted fragment length pattern. This length pattern is different in different individual.
- This is called Restricted Fragment length Polymorphism (RFLP).
4. Southern transfer / Southern blotting : The gel is fragile. It is necessary to remove the DNA from the gel and permanently attaches it to a solid support.
This is accomplished by the process of Southern blotting. The first step is to denature the DNA in the gel which means that the double-stranded restriction fragments are chemically separated into the single stranded form.
The DNA then is transferred by the process of blotting to a sheet of nylon. The nylon acts like an ink blotter and "blots" up the separated DNA fragments, the restriction fragments, invisible at this stage are irreversibly attached to the nylon membrane the "blot".
This process is called Southern blot by the name of Edward Southern (1970).
5. Hybridization : To detect VNTR locus on restricted fragment, we use single stranded Radioactive (P32) DNA probe which have the base pair sequences complimentary to the DNA sequences at the VNTR locus. Commonly we use a combination of at least 4 to 6 separate DNA probes.
Labelled Probes are attached with the VNTR loci of restricted DNA Fragments, this process is called Hybridization.
6. Autoradiography : Nylon membrane containing radio active probe exposed to X-ray. Specific bands appear on X-ray film. These bands are the areas where the radioactive probe bind with the VNTR.
These allow analyzer to identify a particular person DNA, the occurance and frequency of a particular genetic pattern contained in this X-ray film. These x-ray film called DNA signature of a person which is specific for each individual.
The probability of two unrelated individual having same pattern of location and repeat number of minisatellite (VNTR) is one in ten billion (world population 6.1 billion) In India the centre for DNA finger printing and diagnosis (CDFD - center for DNA finger printing & diagnosis) located at Hyderabad.
Application of DNA Finger printing
1. Paternity tests. The major application of DNA finger printing is in determining family relationships. For identifying the true (biological) father, DNA samples of Child, mother and possible fathers are taken and their DNA finger prints are obtained. The prints of child DNA match to the prints of biological parents.
2. Identification of the criminal. DNA finger printing has now become useful technique in forensic (crime detecting) science, specially when serious crimes such as murders and rapes are involved. For identifying a criminal, the DNA fingerprints of the suspects from blood or hair or semen picked up from the scene of crime are prepared and compared. The DNA fingerprint of the person matching the one obtained from sample collected from scene of crime can give a clue to the actual criminal.
CLONING
Clone is the exact carbon copy or copies produced by a single parent (mother or father) by non-sexual methods and are indentical to their parent genetically and morphologically. Clone is a greek word which means twig (Klon=twig). As all the branches of a tree are similar in morphology and genetical characteristics, in the same way clones are also similar to one another.
Cloning is the process of producing many identical organisms (clone), generally used to produce new plants with similar characteristics. Microbes produce clones through asexual reproduction. In higher animals, clones are produced by nuclear transplantation technique in which the nucleus from a somatic cell is transferred into an unfertilized enucleated egg. The world's most famous sheep 'Dolly' was a clone produced by this method.
Many plant species show vegetative reproduction. In these plants, the clones produced by a twig (detached shoot) are similar in their genotype as well as in phenotype (except environmental variations). Scientists have been much curious to apply this characteristics of plants on animals also to conserve the desired genotypes of some rare animals by making their clones. In higher animal, showing sexual reproduction, a zygote is formed after fertilization of the egg by speromatozoan. Zygote differs from its parents in genotype. It was revealed by the scientists through several experiments that only the egg and/or zygote has the potential to produce a whole individual from a single cell.J.B. Gurdon(1969) of Oxford University applied this fact while performing an experiment on frog. He destroyed the nucleus of an unfertilized egg of frog by treating with U.V rays and transferred the nucleus of intestinal epithelial cell of tadpole into the egg cell . In this experiment a few of the many transplanted eggs could develop into tadpoles. These developed tadpoles were identical in genotype and phenotype to their parents. This nuclear transplantation technique devised by Gurdon is still being used in cloning paractice in some modified manner. A brief introductory history of cloning is given in table.
In addition to the fact depicted in table attempts are continuously in progress in this field. In December, 2001 (report published in February, 2002) scientists at University, Texas, successfully produced the first cloned domestic pet named as copy cat (C.C). Further, in Aug. 2005 Woo-Sukhwang of South Korea produced the clone of an Afganian hound (Domestic dog used for hunting).
| An introductory story of cloning | |||
| Year | Name of the Scientist | Brief account of the experiment (s) | Result |
| 1950s | Briggs & King | Nuclear transplantation from embryo to egg in frog. | Tadpoles produced but died before adulthood. |
| 1960s | John B. Gurdon | Nuclear transplantation from cells of skin, liver, kidney into the egg in frog. | Tadpoles produced but died before adulthood. |
| 1970s | Illmense | Nuclear transplantation from embryo to egg in Drosophila | Larvae produced but died before adulthood |
| 1984 | Mcgrath & Solter | Nuclear transplantation from embryo to egg in mouse | A few mice born but none lived to adulthood. |
| 1993 | Hall & Stillman | Artificial splitting of an embryo of human into two identical twins | First artificially twinned embryos developed but abnormal |
| March, 1995 | Roslin Institute team,Scotland | Nuclear transplantation from embryo to egg in sheep | Megan and Moragn sheep born normally |
| Feb, 1997 | Roslin Institute team, Scotland | Nuclear transplantation udder cell to egg in sheep | "Dolly" sheep born normally |
| March, 1997 | Don Wolf and Coworkers, Oregon | Nuclear transplantation from embryo to egg in monkey | Two monkeys "Neti"and "Ditto" born normally |
| Dec. 1997 | Roslin Institute team, Scotland | Nuclear transplantation from embryo to egg in sheep | Molly and Polly sheep born normally |
| 1998 | University of Hawai | Nuclear transplantation from adult cell to egg in mice | 50 mice born normally |
| 1999 | Kato and Coworkers | Nuclear transplantation from adult cell to egg in cow | "George and Charlie" cows born normally |
| 2000 | Well and his Associated | Nuclear transplantation from skin cell to egg in cow | Many cows born normally |
| 2001 | Kabota | Nuclear transplantation from skin fibroblast culture to egg in cow | Six calves born |
An Italian scientist Dr. S. Antenori is trying hard to produce human clone. It has triggered serious discussions and debates at global level, focusing on the medical and ethical issues.
Types of Cloning
Cloning is an extensive technique, which is divided into following types, on the basis of the experimental material used -
(1) Gene cloning (2) Microbial cloning (3) Cell cloning (4) Plant cloning and (5) Animal cloning
Animal cloning
Embryonic cell in animals, are deprived of their totipotency by the time they enter into gastrula stage. So animal cloning is some what more difficult than plant cloning. On the basis of aims and end products animal cloning is of two types
(i) Reproductive and
(ii) Therapeutic.
A clone of the whole animal is prepared in reproductive cloning. The techniques which are used in such cloning include blastomere separation, nuclear transplantation and the Honolulu technique. This technique can be used to conserve and increase the number of those animals which are threatened to be extinct in the near future.
In nuclear transplantation technique, nucleus is removed from the egg obtained from the female. After then the nucleus of the desired cell (2n) is transferred into the enucleated egg cell which is then allowed to develop into an embryo in suitable conditions. The developing embryo is the clone of the donor cell from which nucleus was obtained for transplantation.
In Honolulu technique ( 1998-Teruhiko wakayama - cloned mice) there is no fusion of the donor and recipient cells or its nucleus. Instead of ,nucleus from the donor cell which is in G0 or G1, stage is substituted into enucleated egg cell (culture medium or chemical both is used in place of electric shock to stimulate development).
The rapeutic cloning technique may prove to be very useful in the field of medical science, particularly when there is a need for the transplantation to replace some damaged and diseased organ or tissue by a healthy organ from a suitable donor. In such condition, if the cells from the patient himself are taken and cultured to form desired organ. The organs developed in such manner( organ cloning) will be easily acceptable by the patient, and there will be no possibility of its rejection as often occurs otherwise.
A number of diseases like will be no possibility of its rejection as often occurs otherwise . A number of diseases like parkinsonia, alzheimer, diabetes and diseases related with kidneys will possibly be cured in future by the application of cloning technique. "Dolly" sheep was produced by using nuclear transfer technique by Dr. lan wilmut and his collegues at Roslin Institute of Scotland in 1997.
They used somatic cells from udder (mammary glands) for forming this clone. One udder cell with its nucleus intact was selected because this nucleus carried the mother's genetic information.
Meanwhile, and unfertilized egg cell was taken from a different sheep. Its nucleus was sucked out and an enucleated egg cell was obtained. After then the udder cell nucleus was fused with the enucleated egg cell under electrical stimulation. Now this egg cell had the mother's nucleus. At last the fused egg was implanted into the uterus of surrogate mother, other than the egg donor where it grew into a lamb. Thus the Dolly was born, as a genetically identical copy of its mother


Advantages of Cloning
1. This technique can be used to improve the breeds of live-stock used in agriculture.
2. Cloning is helpful for providing many useful substances and chemicals required for human body, and also in the cloning of such animals which can be used as a source of organs for transplantation purpose in medical practices.
3. British scientists have been successful in obtaining alpha antitrypsin from sheep for curing an incurable disease emphysema and clotting factor- ix for curing haemophilia. It has been possible by the use of genetic engineering.
4. Many incurable diseases which are not curable so far, may be cured effectively in near future by applying the processes relating with cloning and genetic engineering.
5. Cloning is useful to increase the number of individuals of those species which are at the edge of extinction, thus helpful in conservation of biodiversity.
Disputes Related to Cloning
At present, the issue of cloning has been a matter of discussion and disputes among the scientists and the sociologists. Many questions are being raised with regards to the ethical, moral, and social aspects of cloning. Doubts have also been there about the health and ageing of clones and the misuse of cloning.
Cloning has important role in treatment of serious and non-curable diseases, a view that is favoured by most scientists, however there is no agreement on the issue of human cloning.
|
26 videos|287 docs|64 tests
|
FAQs on Genetics - Additional Study Material for NEET
| 1. What is NEET and how does it relate to genetics? |  |
| 2. What are the important topics in genetics for the NEET exam? |  |
| 3. How can I prepare effectively for the genetics section of the NEET exam? |  |
| 4. Are there any specific genetic disorders that are frequently asked in the NEET exam? |  |
| 5. Can you provide some tips for answering genetics-related questions in the NEET exam? |  |

|
Explore Courses for NEET exam
|

|


















